RIC-seq
RIC-seq
RNA In situ conformation sequencing
RNA in situ conformation sequencing (RIC-seq) is a novel technology for capturing the higher-order structures of RNA and identifying molecular interaction sites at the cellular level. RIC-seq can investigate the conformation and organizational rules of mRNA and non-coding RNA in cells, map enhancer-promoter regulatory networks, and elucidate novel mechanisms of enhancer-mediated oncogene activation. Additionally, RIC-seq can also be used to systematically analyze the effects of disease-associated mutations on RNA higher-order structures and target interactions, which may reveal the pathogenic mechanisms of mutations in non-coding regions and lay the foundation for clinical diagnosis and treatment.
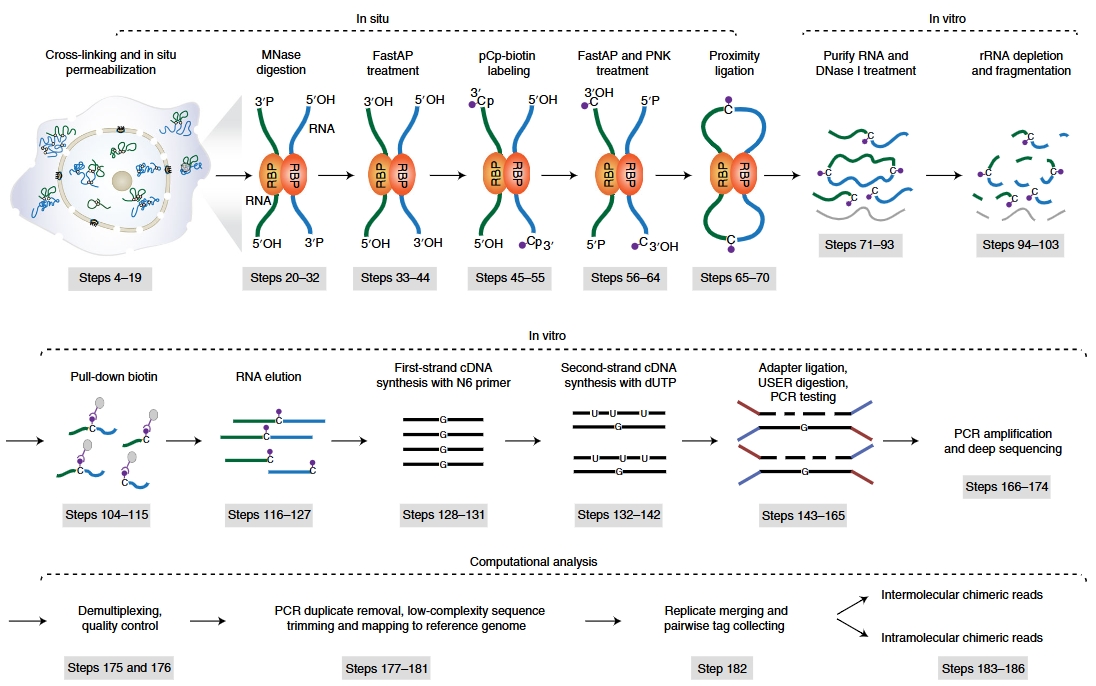
Workflow of RIC-seq (Cao C,et al. Nat Protoc. 2021.)
Applications
1. Identify nuclear RNA–RNA interactions
2. Systematically analyze nuclear non-coding RNA structure and targets
3. Map high-order RNA structures
4. Construct RNA spatial interaction networks
5. Reveal pathogenic mechanisms of non-coding region mutations
Basic Analysis
1. Data quality control
2. Removal of duplicate sequences
3. RNA ribosomal sequence alignment
4. RNA sequence genomic alignment
5. RNA interaction pairing
6. IGV visualization of interactions
Advanced Analysis
1. Global RNA-RNA interaction analysis
2. RNA-RNA topological domain analysis
3. RNA secondary structure analysis
4. RNA functional structure analysis
5. Identification of hub RNAs
6. Enhancer-Promoter interaction network analysis
7. SuperEnhancer-Promoter interaction functional analysis
8. Promoter–promoter interaction functional analysis
Demo
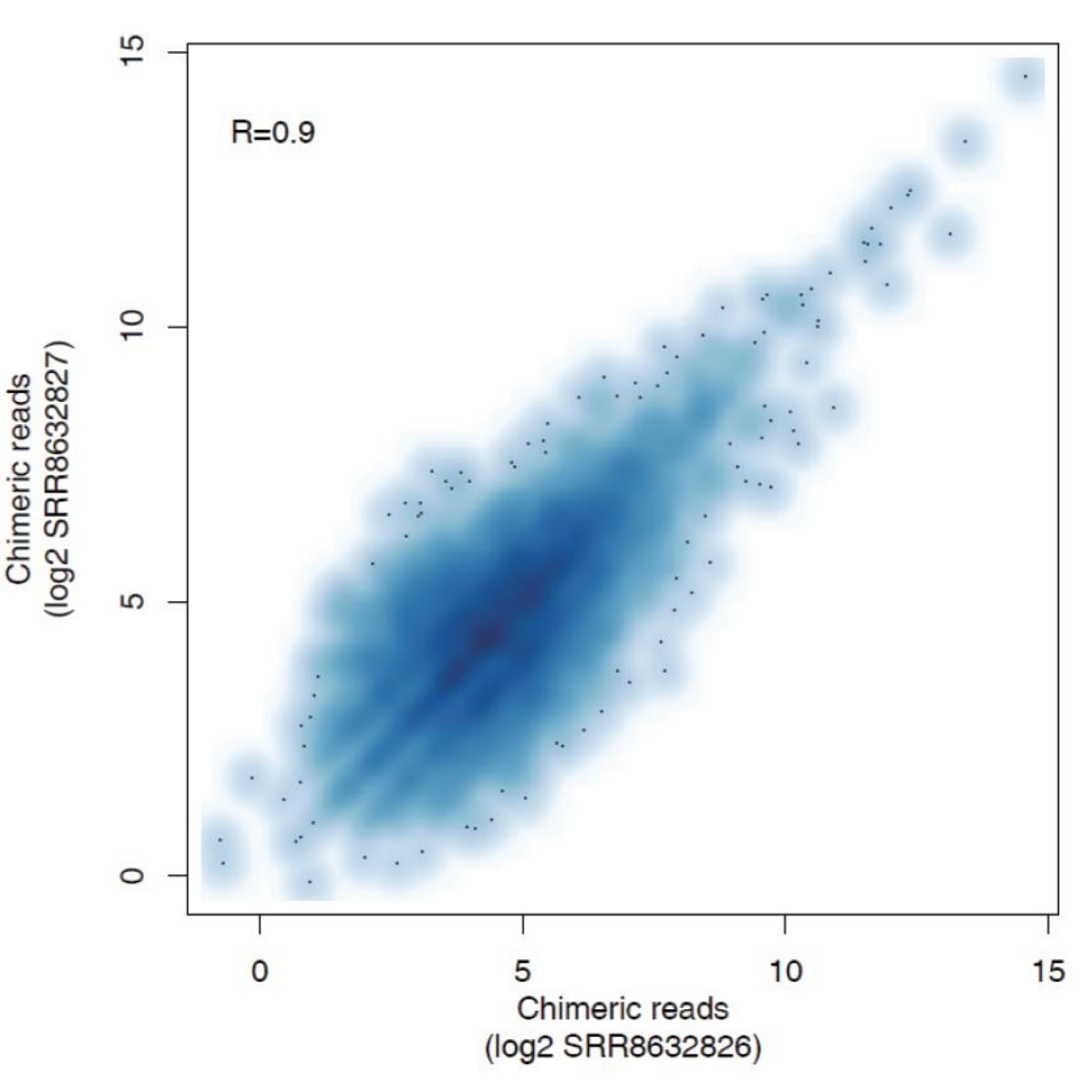
Sampe Correation Matrix
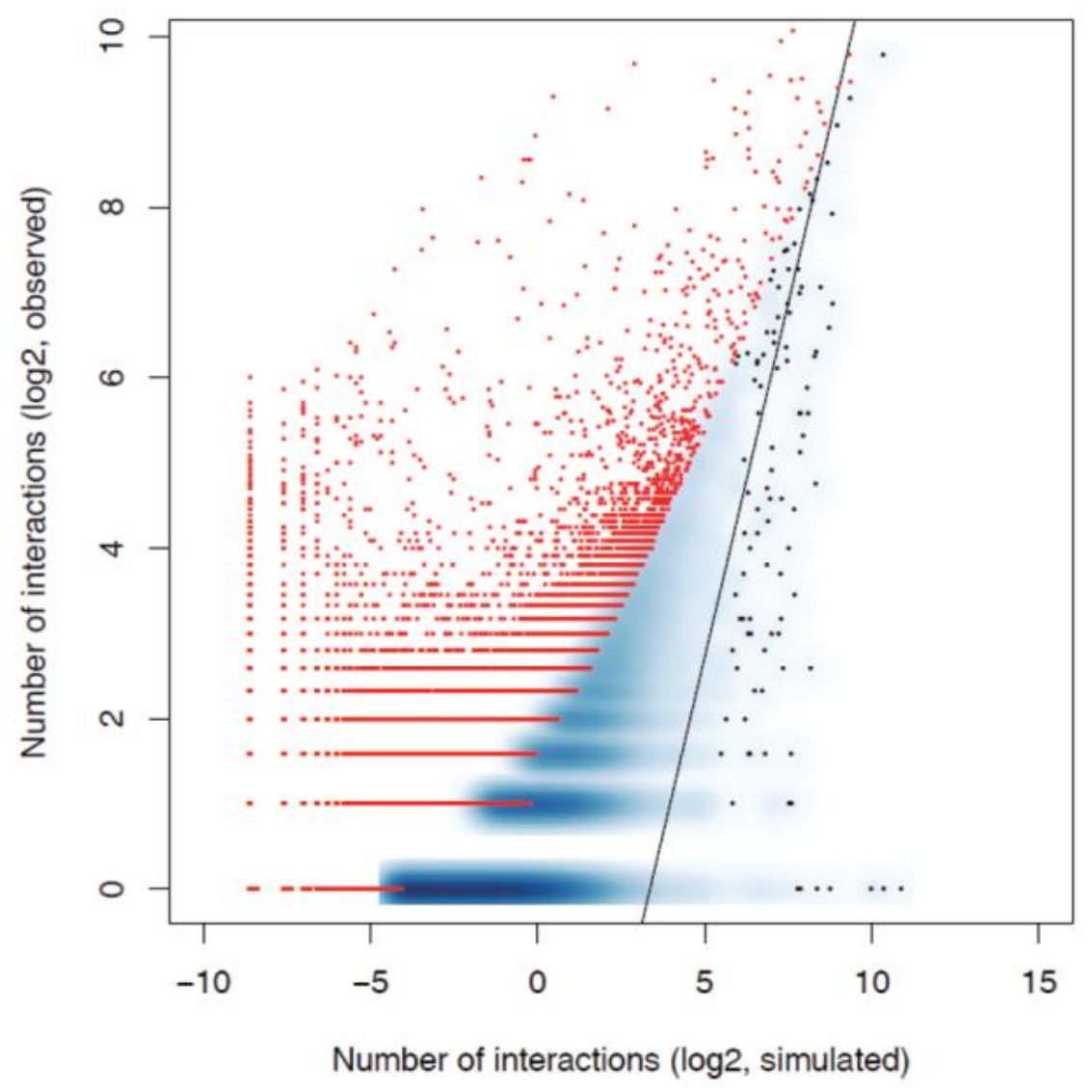
Disribution map of high Confident interacion molecules
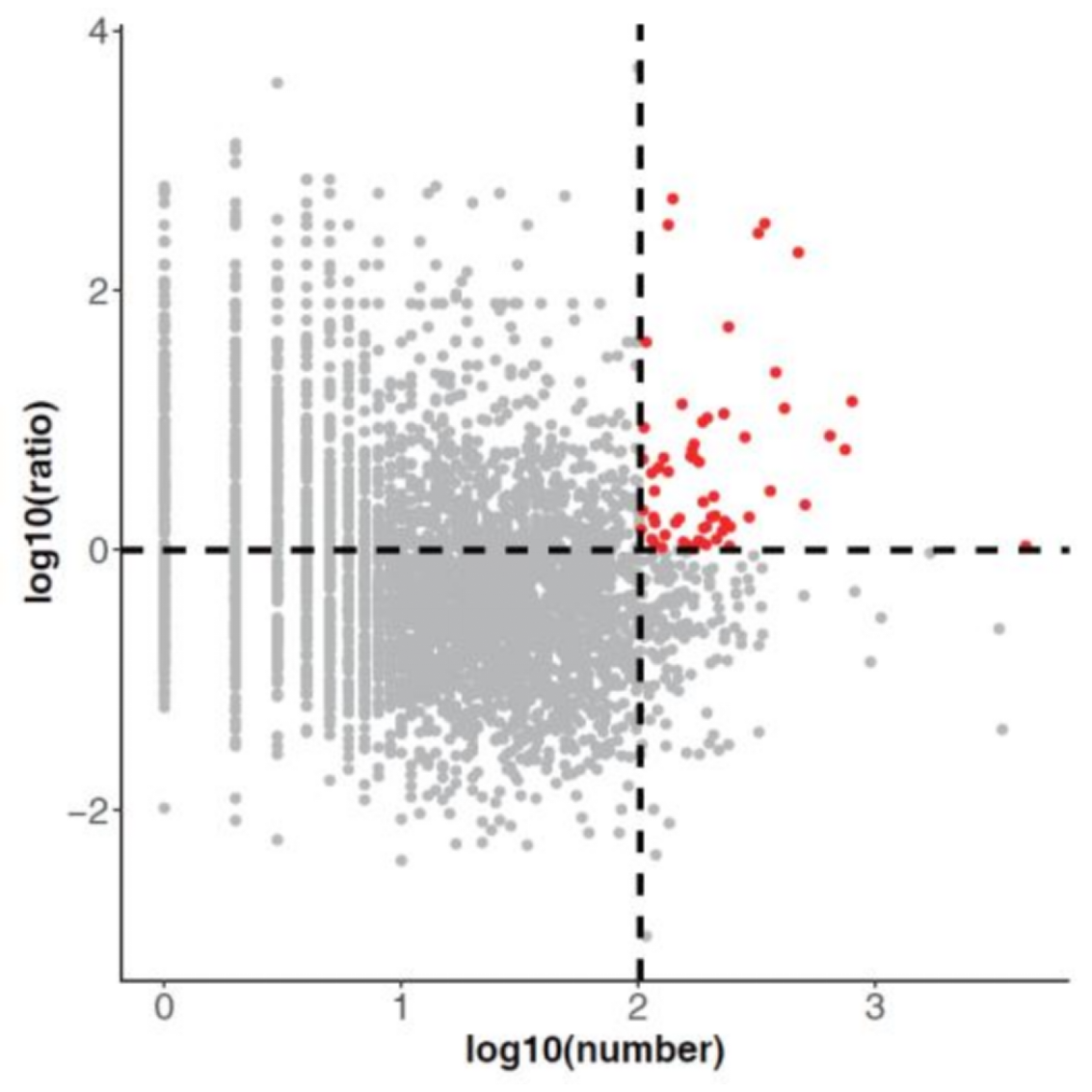
Identification of hub RNAs
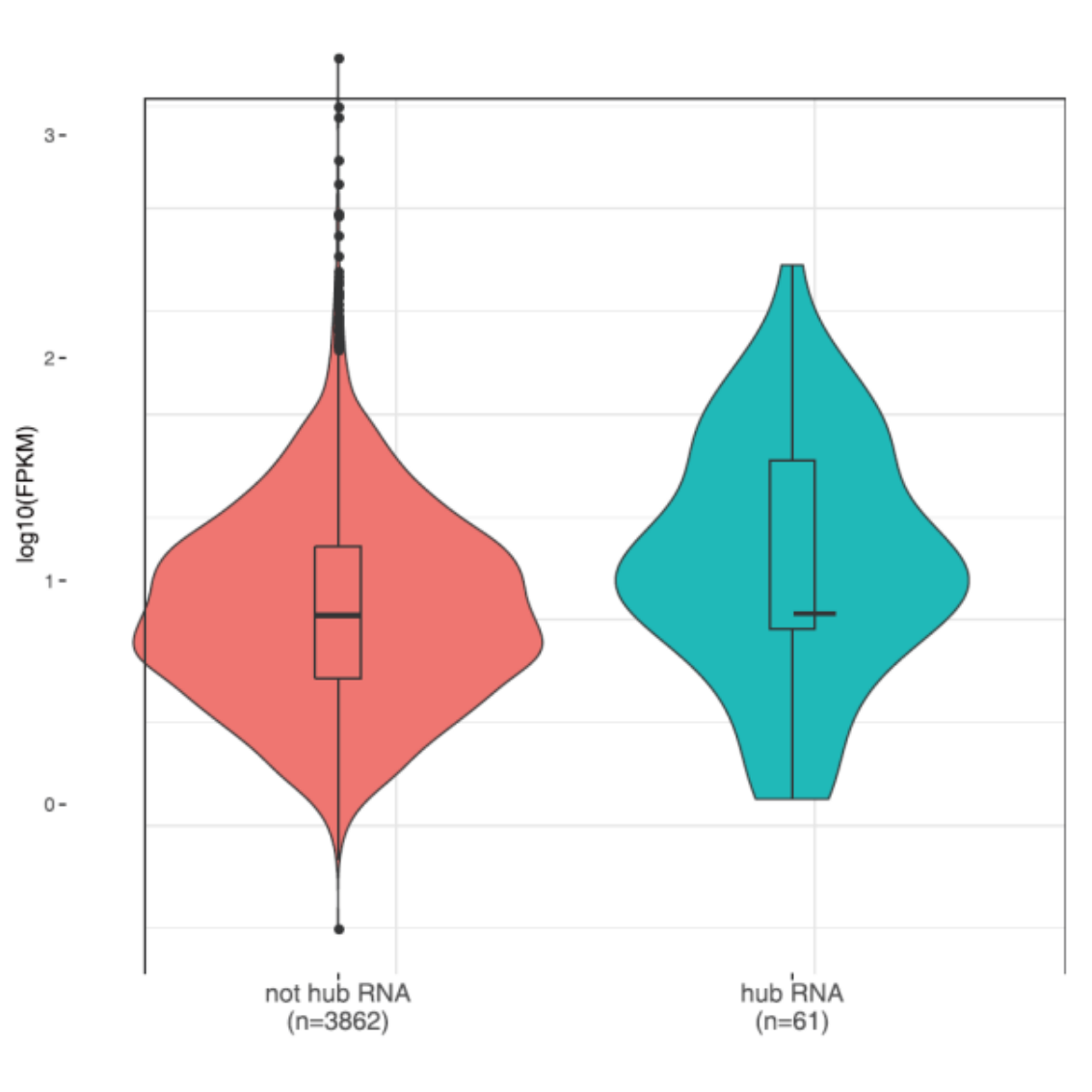
Violin plot of hub RNAs and RNA expression levels
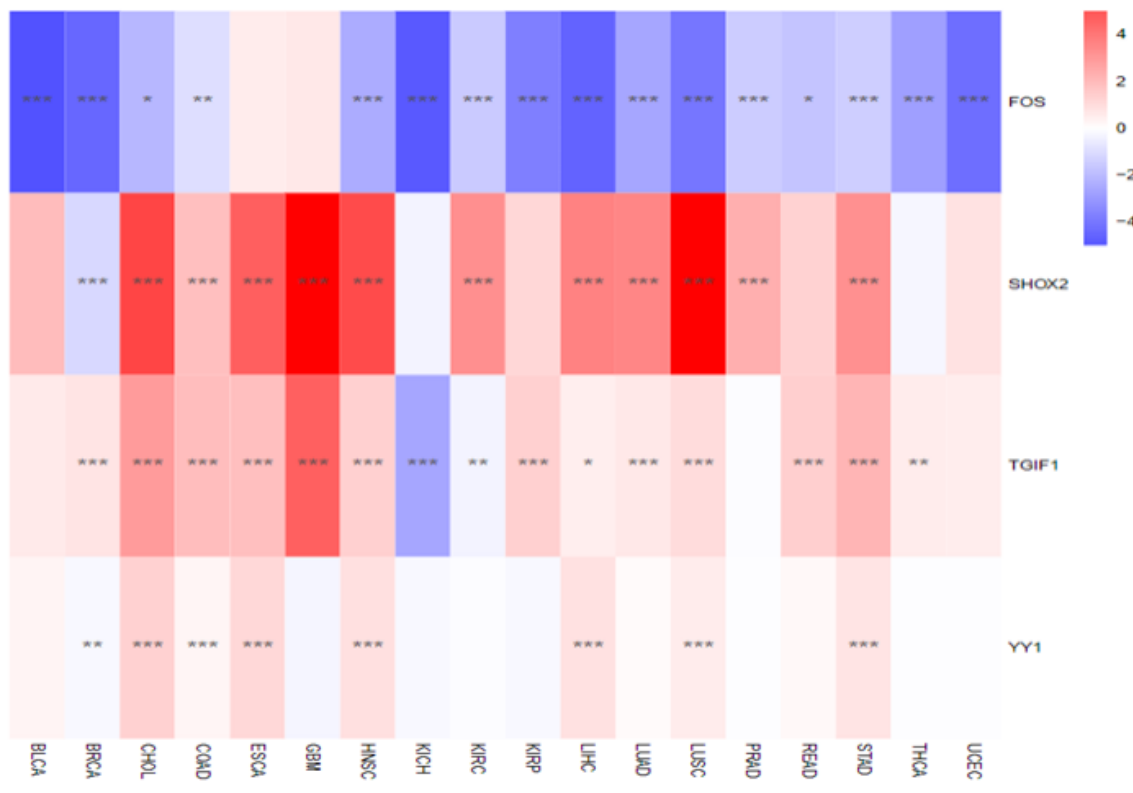
eRNA–uaRNA interaction heatmap within core regulatory circuits
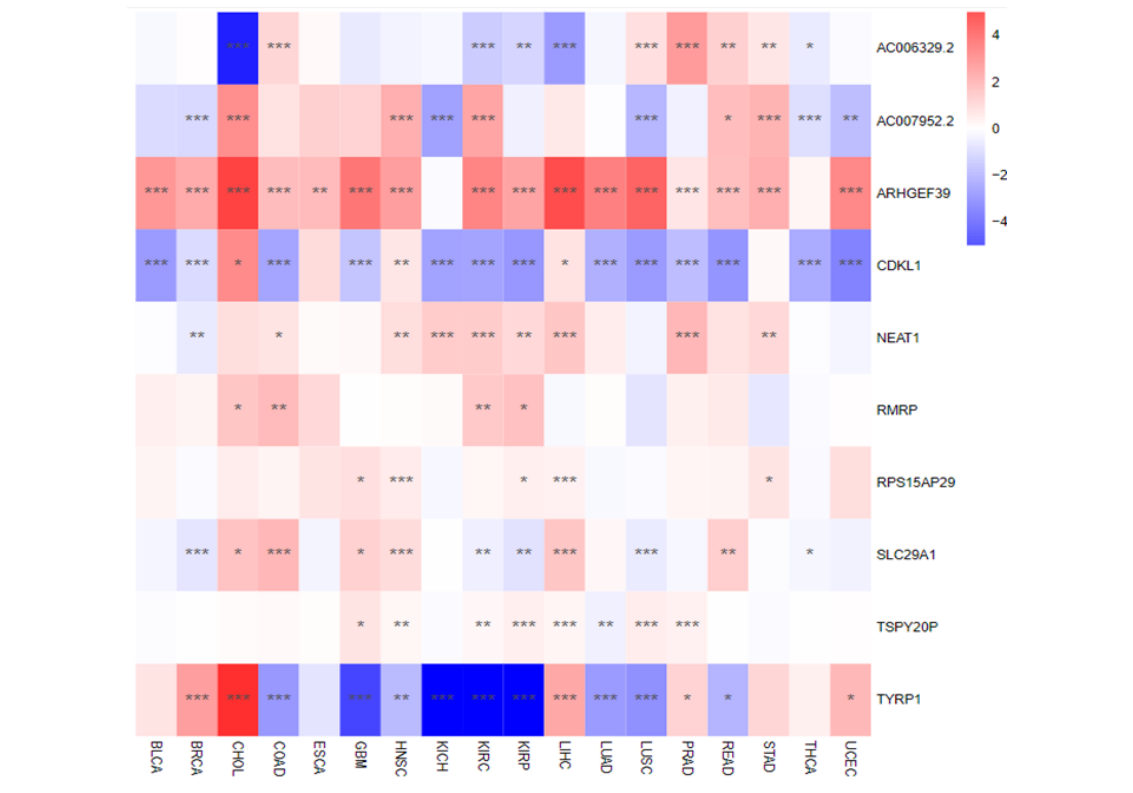
Expression profile changes of ua-hubRNA across 18 common cancers in TCGA database
EPIBIOTEK® Project Publications
1. Lu B, Chen S, Guan X, et al. Lactate accumulation induces H4K12la to activate super-enhancer-driven RAD23A expression and promote niraparib resistance in ovarian cancer. Mol Cancer. 2025;24(1):83. Published 2025 Mar 19.
2. Wang JY, Zhang XP, Zhou HK, et al. The selective sponging of miRNAs by OIP5-AS1 regulates metabolic reprogramming of pyruvate in adenoma-carcinoma transition of human colorectal cancer. BMC Cancer. 2024;24(1):611. Published 2024 May 21.
