DRUG-seq
DRUG-seq
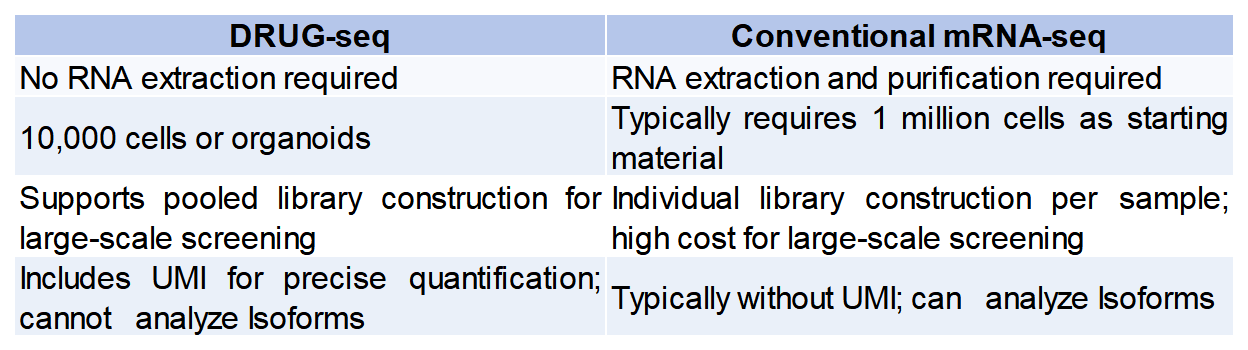
Digital mRNA with pertUrbation of Genes (DRUG-seq) is a rapid transcriptomic sequencing technology designed for low-input cell and tissue samples. The method utilizes oligo-dT magnetic beads carrying Barcode tags and Unique Molecular Identifiers (UMIs) to efficiently bind to the Poly(A) tail of mRNA for reverse transcription and library construction.
The barcoded tags on the magnetic beads allow for the pooling of multiple samples into a single library, while the UMI on each sequence enables precise quantification of captured mRNA molecules, minimizing data bias caused by PCR amplification. DRUG-seq significantly reduces sample input requirements and sequencing costs, making it widely applicable in basic life science research, translational medicine, and new drug development.
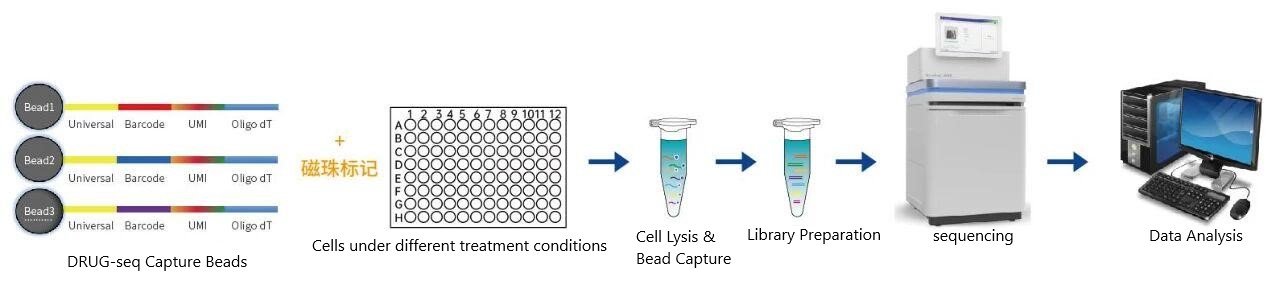
Workflow
Applications
1. High-Throughput Profiling: Transcriptomic mapping of FACS-sorted cells.
2. Functional Genomics: High-throughput gene function studies.
3. Target Identification: Analysis of compound mechanisms of action and targets.
4. Drug Screening: Screening for tumor-targeted therapies.
5. Drug Discovery: Development of novel pharmaceutical agents.
Sample Requirements
1. Type: Cells
2. Input: 1×10^3-4 cells/sample
3. Cell Viability: ≥85%
4. Species Compatibility: Validated for Human, Mouse, and Rat. (Other species require a feasibility assessment).
Bioinformatics Analysis
1. Data Quality Control
2. Gene Alignment & Statistics
3. Gene Expression Quantification
4. Differential Expression Analysis
·Heatmap of Differentially Expressed Genes
·Volcano Plots
5. Gene Ontology (GO) Functional Analysis
6. KEGG Pathway Analysis
7. Gene Set Enrichment Analysis (GSEA)
Demo
1. He X, Wang X, Wang H, et al. NeuroD1 Regulated Endothelial Gene Expression to Modulate Transduction of AAV-PHP.eB and Recovery Progress after Ischemic Stroke. Aging Dis. 2023;15(6):2632-2649. Published 2023 Dec 20. doi:10.14336/AD.2023.1213
2. Chen T, Liu S, Yang Z, et al. Investigation roles of Adamts1 and Adamts5 in scleral fibroblasts under hypoxia and mice with form-deprived myopia. Exp Eye Res. 2024;247:110026. doi:10.1016/j.exer.2024.110026
3. Liu L, Ha S, Cao D, Li M, Li Z. Transposition element MERVL regulates DNA demethylation through TET3 in oxidative-damaged mouse preimplantation embryos. Mol Med. 2025;31(1):95. Published 2025 Mar 12. doi:10.1186/s10020-025-01143-3
