GRO-seq
GRO-seq
Global nuclear Run-On sequencing (GRO-seq) is a specialized method designed to measure nascent RNA (Core L J, et al. Science, 2008.), enabling the systematic identification of enhancer RNAs (eRNAs). The protocol involves the rapid cryopreservation of cells to arrest transcriptional complexes within the nucleus. Following nuclei isolation, a "run-on" buffer is introduced to resume RNA transcription in the presence of Br-UTP. The resulting BrU-labeled nascent RNA is immunopurified using anti-BrdU antibodies and subjected to high-throughput sequencing.
Because changes in nascent RNA directly reflect immediate transcriptional regulation mechanisms, GRO-seq technology allows for the observation of genome-wide active transcription. It generates high-resolution maps depicting the position, density, and orientation of active RNA Polymerase II.
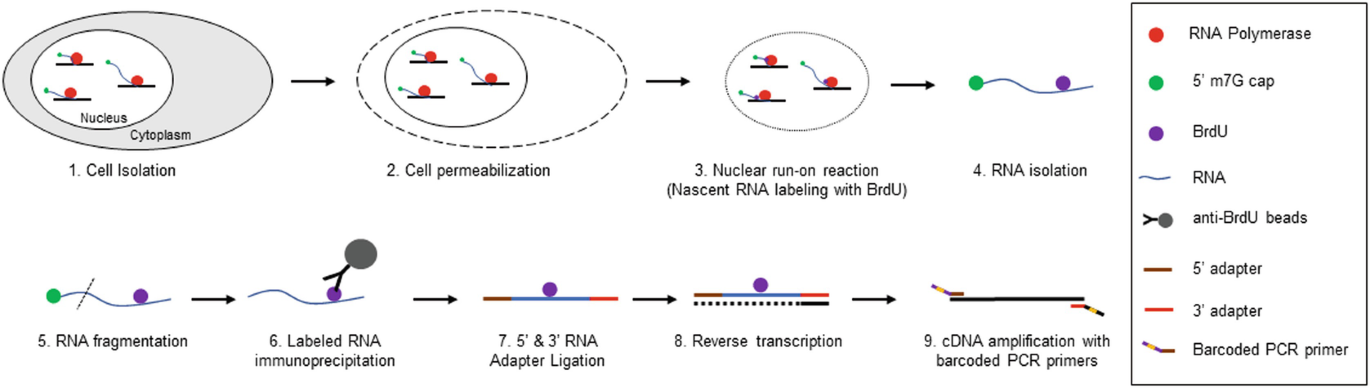
Tzerpos P, et al. Methods Mol Biol. 2021.
Workflow
Sample Requirements
Type & Input
Cells,≥1×10⁷ cells/sample
Validated for Human, Mouse, and Rat. (Other species require a feasibility assessment).
Applications
1. Supersedes conventional RNA-seq by enabling precise detection of gene expression during active transcription;
2. Precisely determines transcription start sites (TSSs) and transcriptional orientation;
3. Enables identification of direct transcriptional targets;
4. Facilitates discovery of novel transcripts, including non-coding RNAs;
5. Enables identification of enhancer RNAs (eRNAs);
Bioinformatics Analysis
1. Data QC: adapter trimming, low-quality read filtering, and contamination removal.
2. Genome alignment and transcriptome-wide read distribution analysis.
3. Meta-analysis of read enrichment at TSSs.
4. Analysis of transcriptional pausing.
5. eRNA identification and analysis.
6. Differential expression analysis.
7. GO/KEGG analysis of differentially expressed genes.
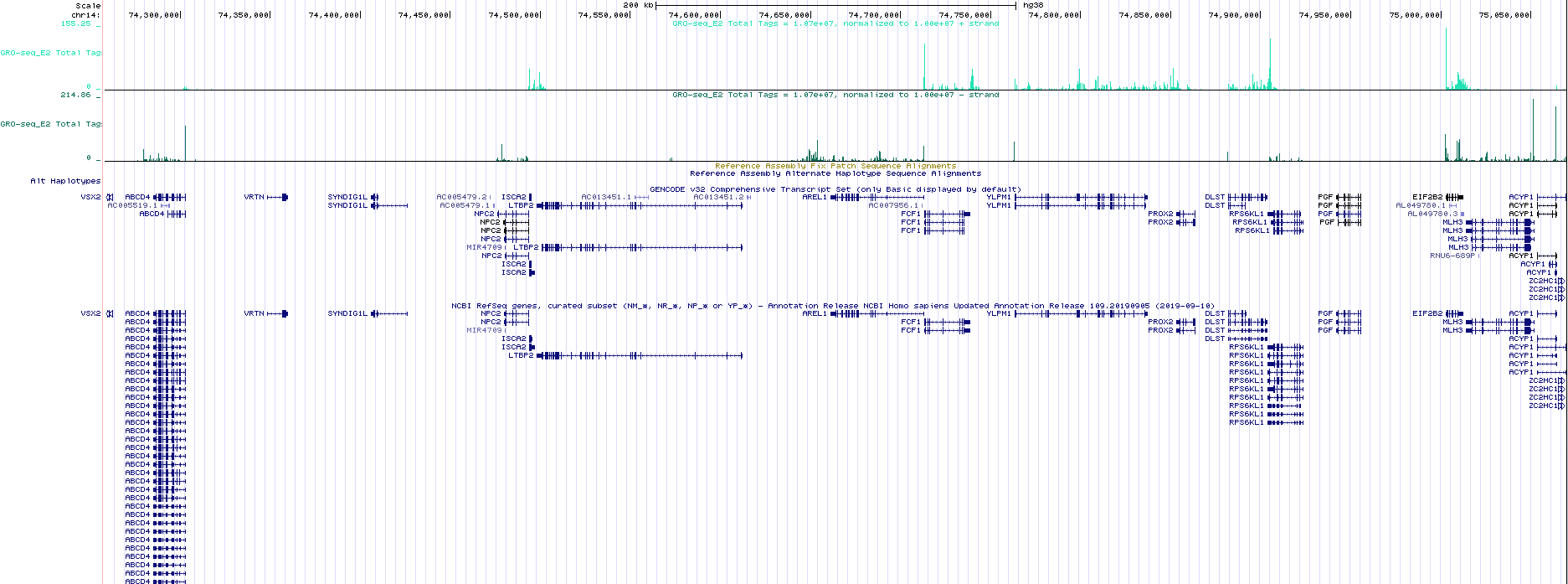
Visualization of GRO-seq data (bed files) in the UCSC Genome Browser
Demo
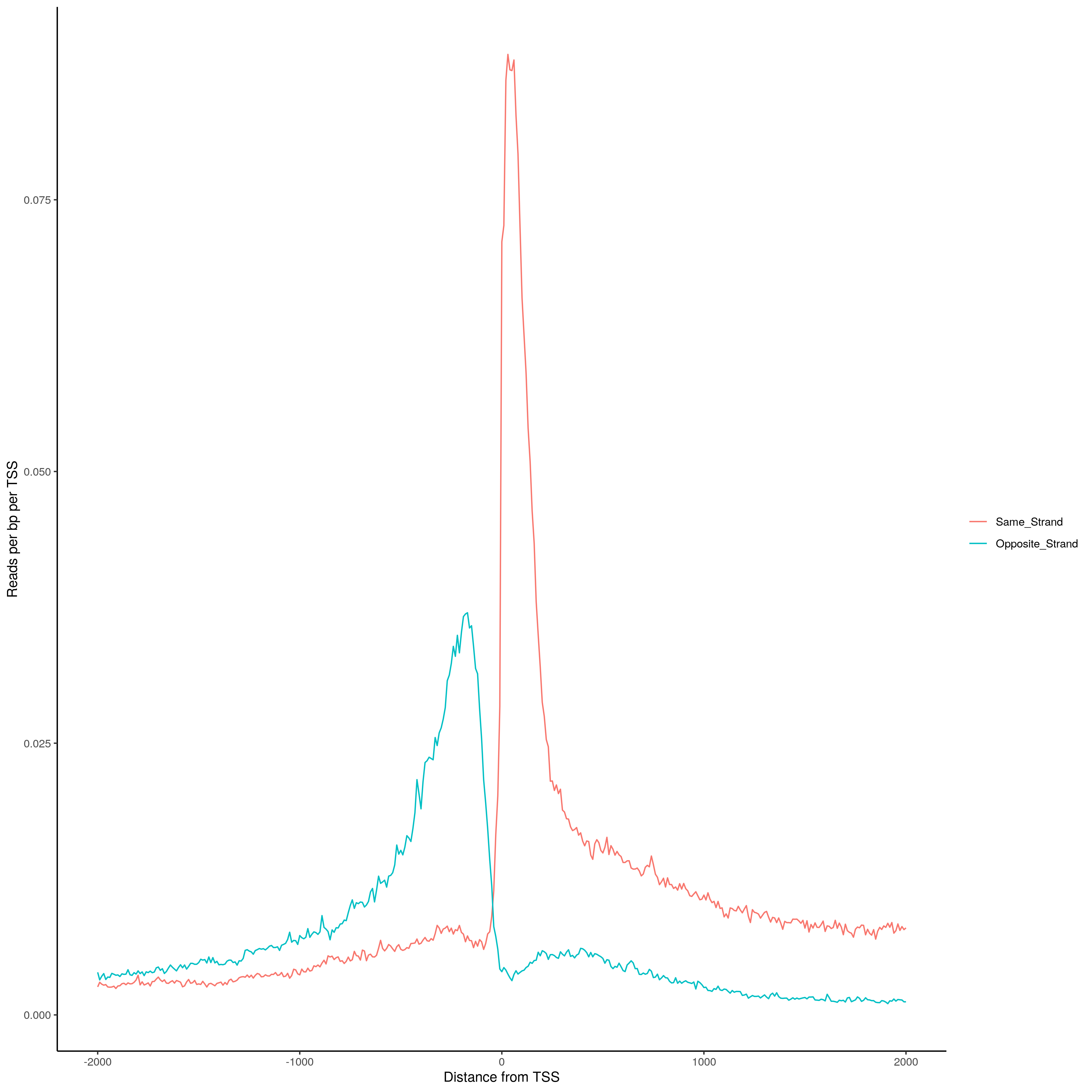
Histogram of TSS distances
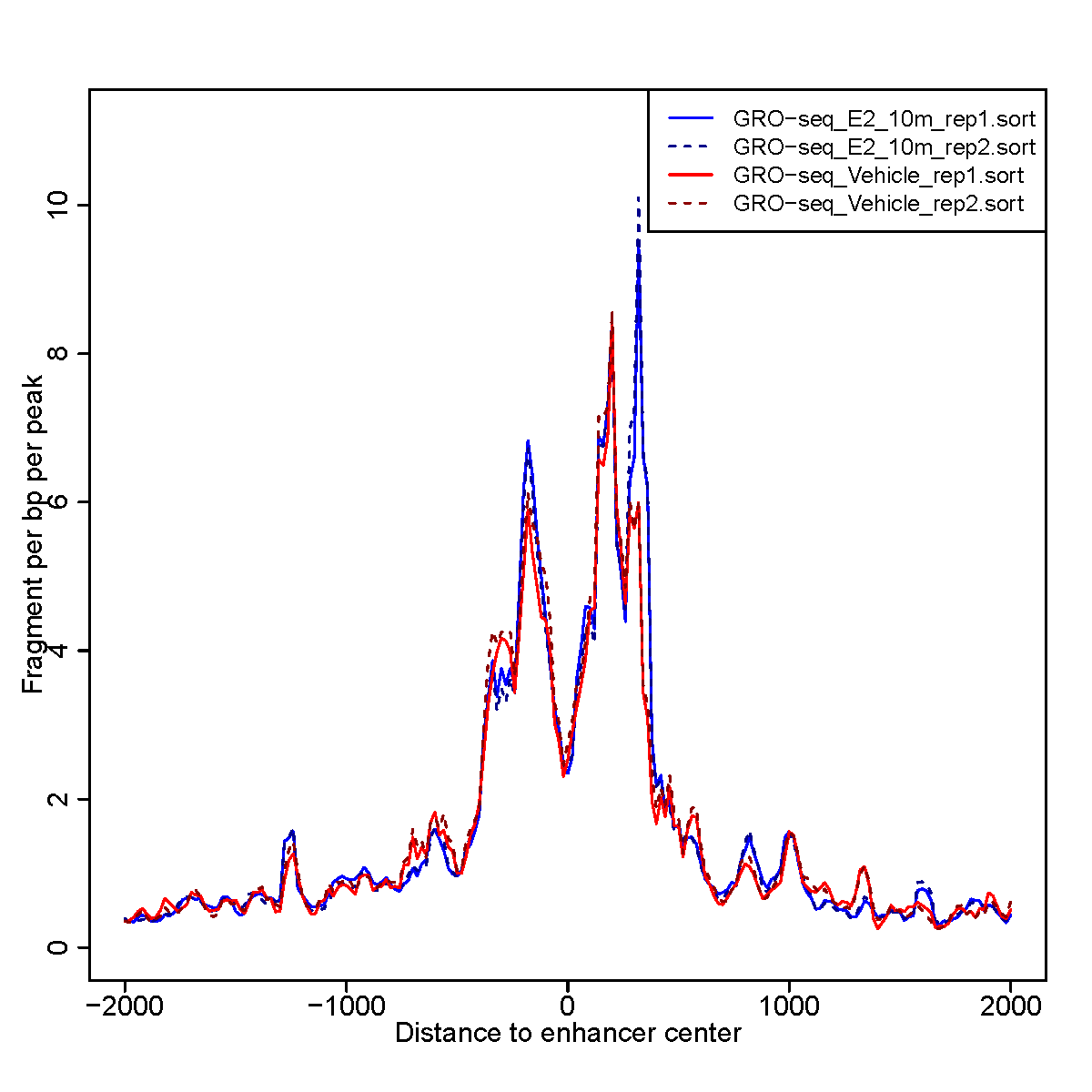
GRO-seq signal enrichment centered on enhancer regions across samples
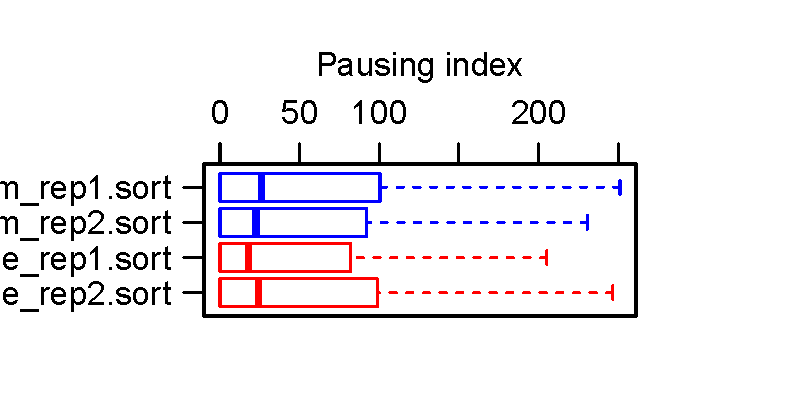
Boxplot illustrating changes in the Pausing Index

Heatmap of condition-dependent transcriptional changes in active genes surrounding the TSS
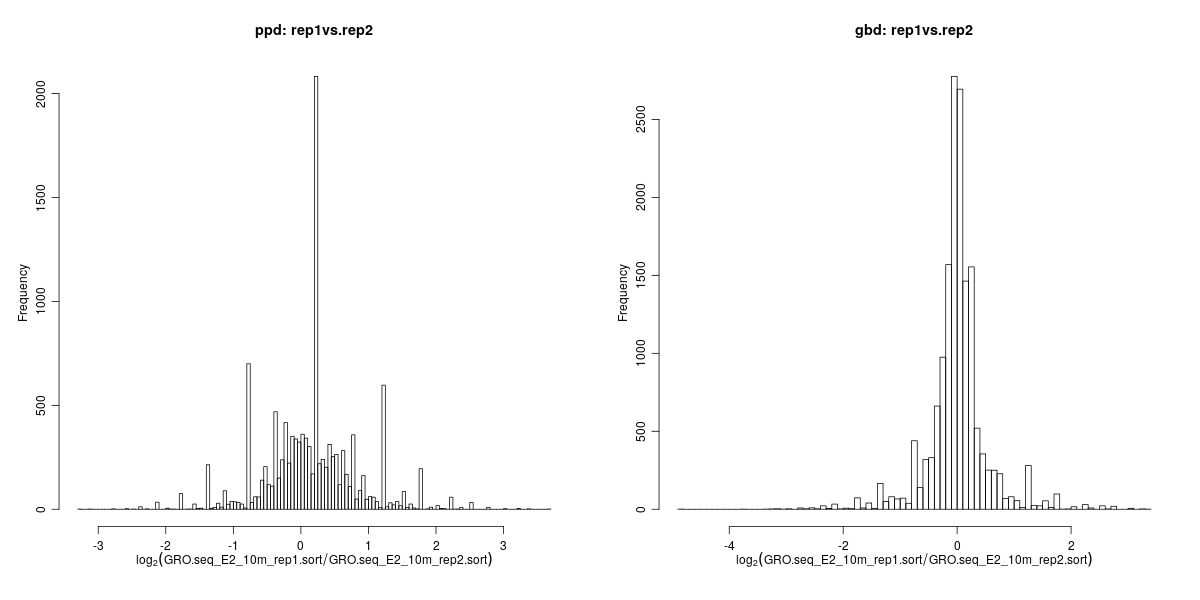
Histogram showing transcriptional variation between replicate samples
EPIBIOTEK® Project Publications
Wang WL, Song ZX, Bu SM, et al. Ythdc1-p300-Klf5 Complex-Mediated Golgi Dysfunction Promotes Aortic Aneurysm. Adv Sci (Weinh). 2026;13(4):e12116. doi:10.1002/advs.202512116
