OmniRNA-seq
OmniRNA-seq
OmniRNA-seq, a whole transcriptome sequencing service, addresses the dependency of traditional library construction methods on RNA terminal modifications, enabling more comprehensive transcriptome coverage and facilitating the construction of multidimensional RNA profiles. OmniRNA-seq features high sensitivity, with its core advantage lying in the detection of a broader spectrum of RNA species and fragments. A single sequencing run enables comprehensive analysis of multiple RNA molecules, including mRNA, lncRNA, and sncRNA (such as miRNA and tsRNA). This technology provides robust technical support for disease biomarker development, early cancer screening, and related research applications.
Advantages
1. High sensitivity for trace samples with low input requirements
2. Broad-spectrum RNA capture capability for efficient library construction with substantially enhanced performance
3. Whole transcriptome coverage (mRNA, lncRNA, miRNA, piRNA, tsRNA, ysRNA, snoRNA, and rsRNA)
Applications
1. Liquid Biopsy & Early Cancer Screening: Detection of circulating RNA markers.
2. Disease Biomarker Discovery: Screening and development of novel diagnostic markers.
3. Intercellular Communication: Investigation of RNA-mediated signaling mechanisms.
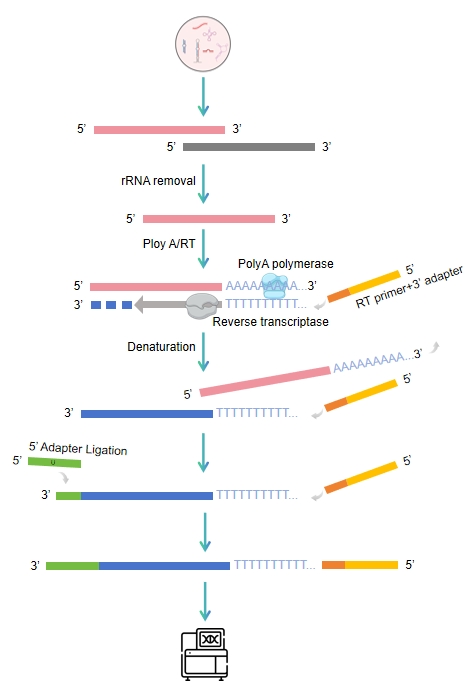
Workflow
Sample Requirements
Limited to human, mouse, and rat. Other species require evaluation
≥ 200 ng
Total RNA
≥ 50 ng
Plasma cfRNA/Exosomal RNA/Cell supernatant RNA
Bioinformatics Analysis
Basic Analysis
1. Raw Data Filtration and Quality Control
2. Reference Genome Alignment
3. Alignment to Various RNA Databases
4. Quantitative Analysis of Various RNAs
5. Differential Expression Analysis of Various RNAs
6. Length Distribution and Classification Annotation of Various sncRNAs
7. Clustering Analysis of Differentially Expressed RNAs
8. GO/KEGG Analysis of mRNAs
Advanced Analysis
1. Customer-Selectable Analysis (for no more than 50 sncRNAs):
(1) Target Gene Prediction
(2) GO Analysis of Target Genes
(3) KEGG Analysis of Target Genes
2. For Multiple Samples, Complimentary Service Available:
Disease Diagnosis / Prognosis Assessment Model Construction (For details, please inquire)
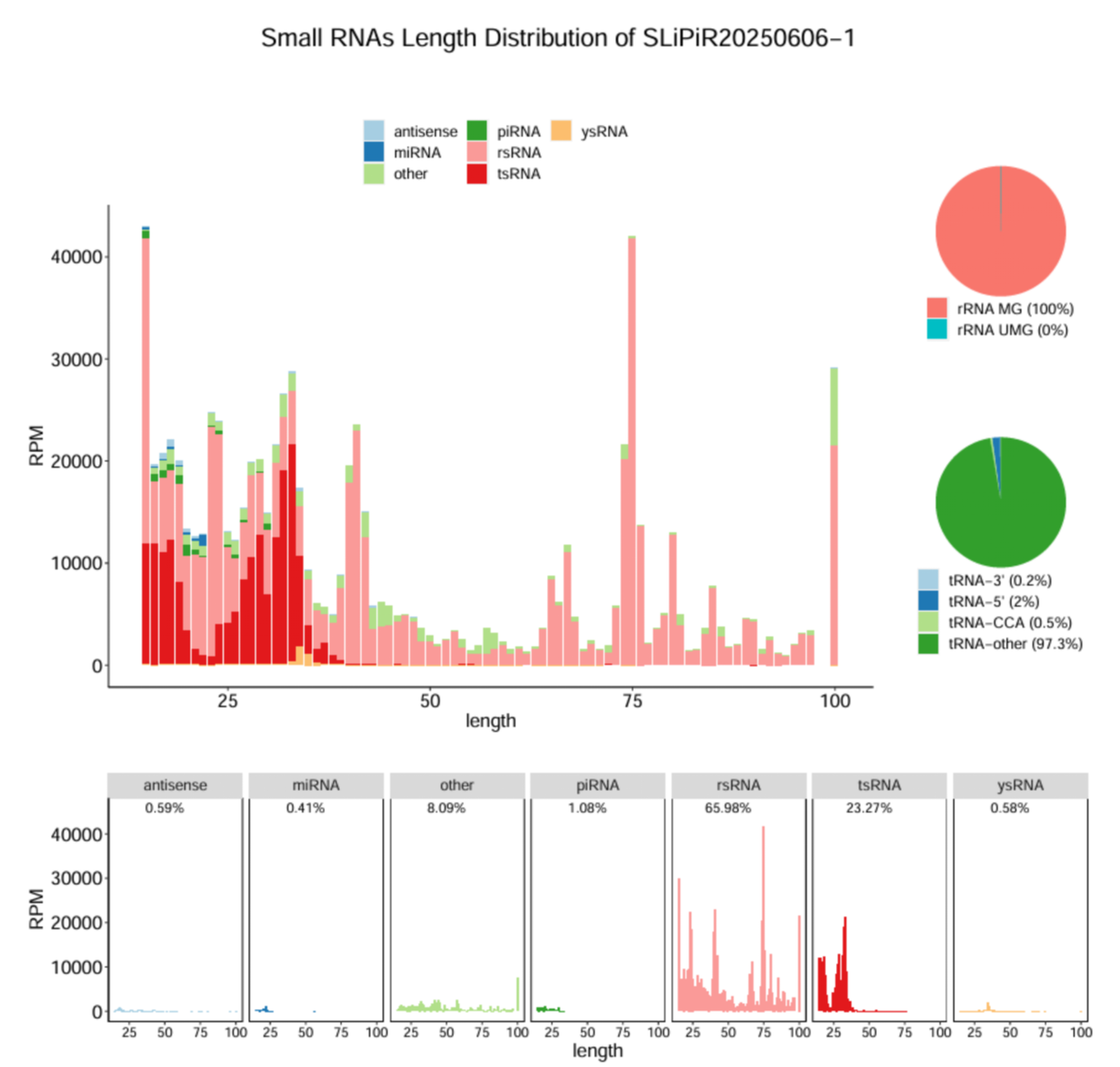
Length distribution and annotation results for sncRNA categories
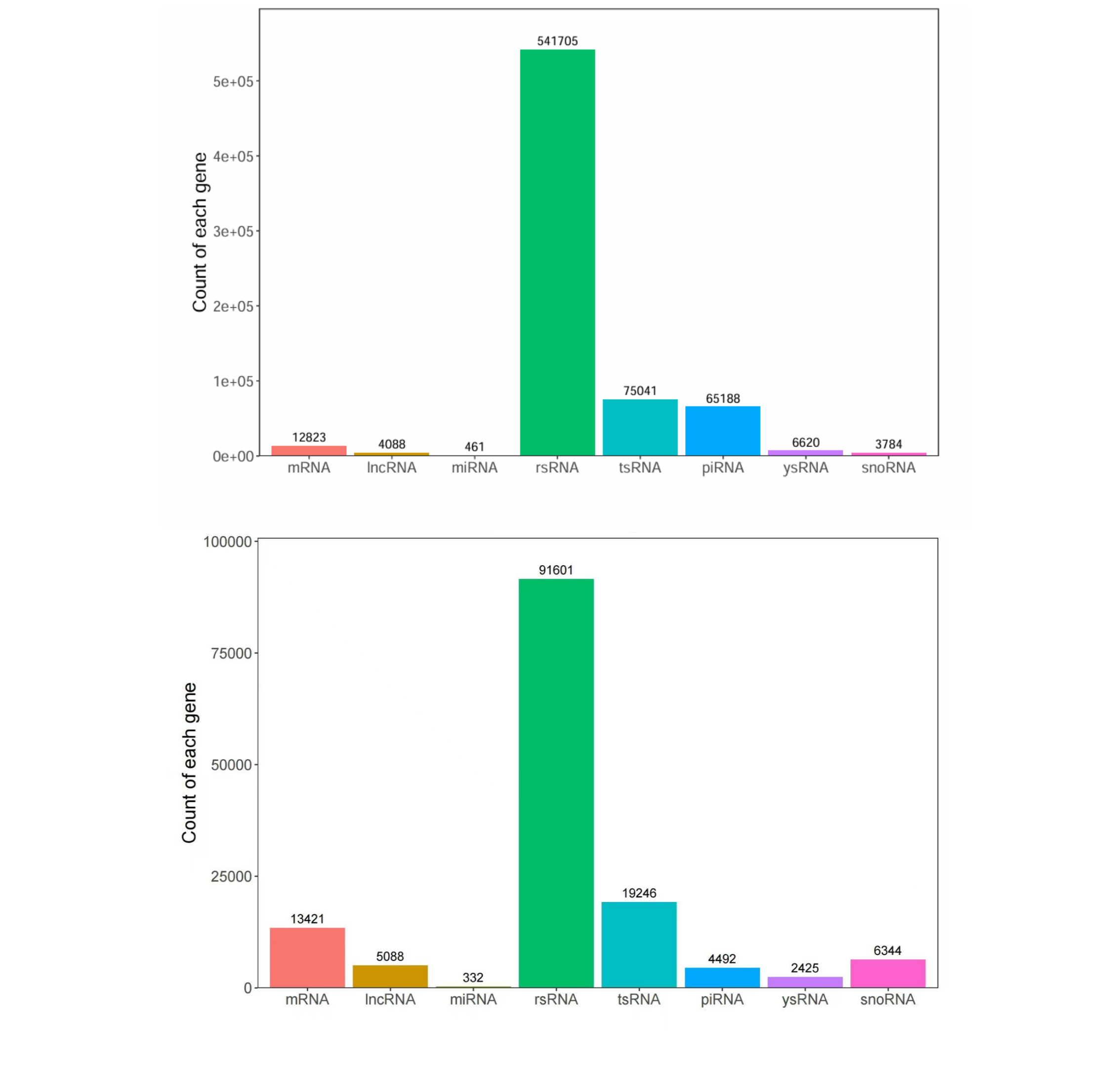
Summary of counts for various RNA categories
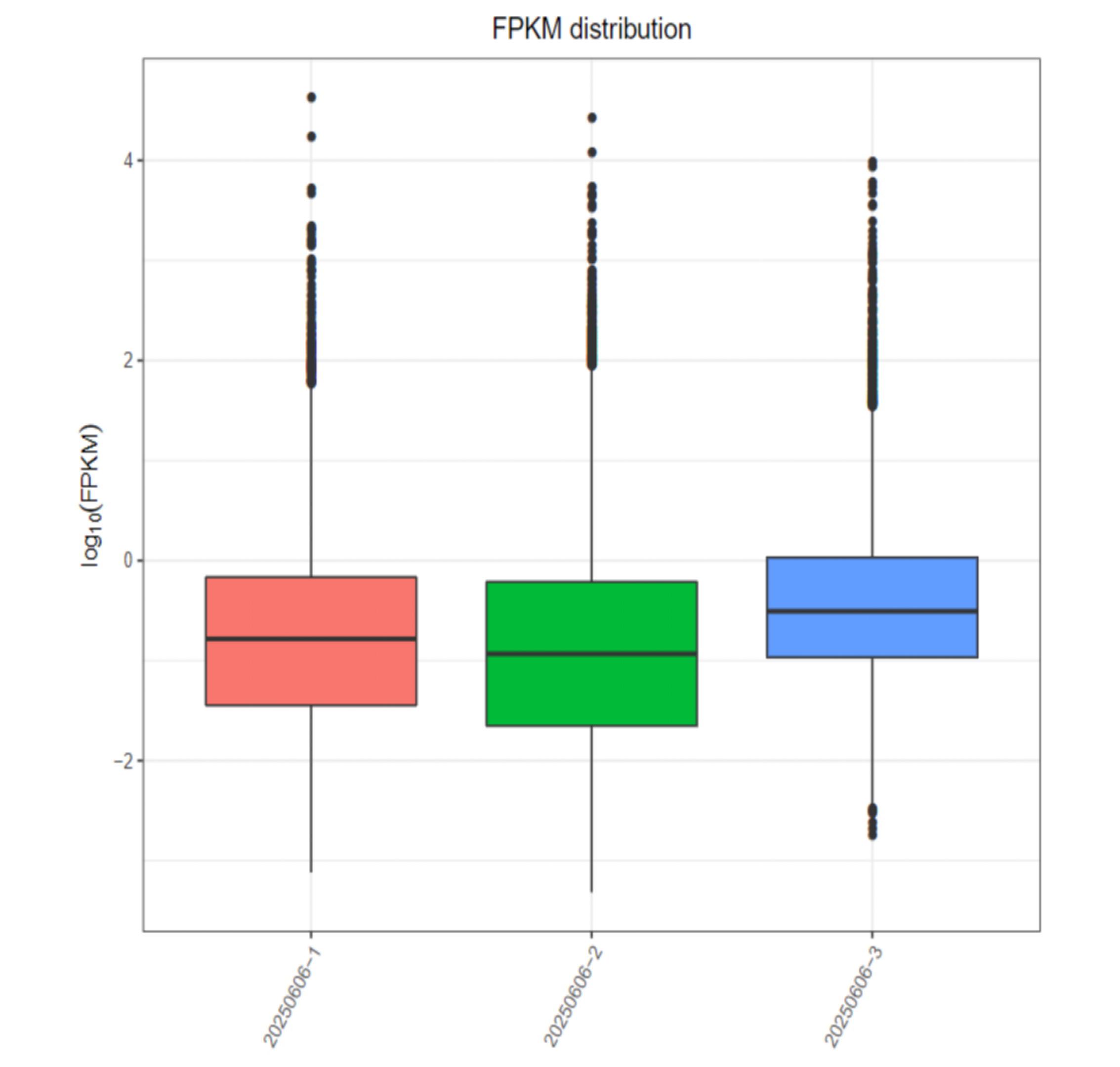
Quantitative analysis results for long RNAs (mRNA, lncRNA)
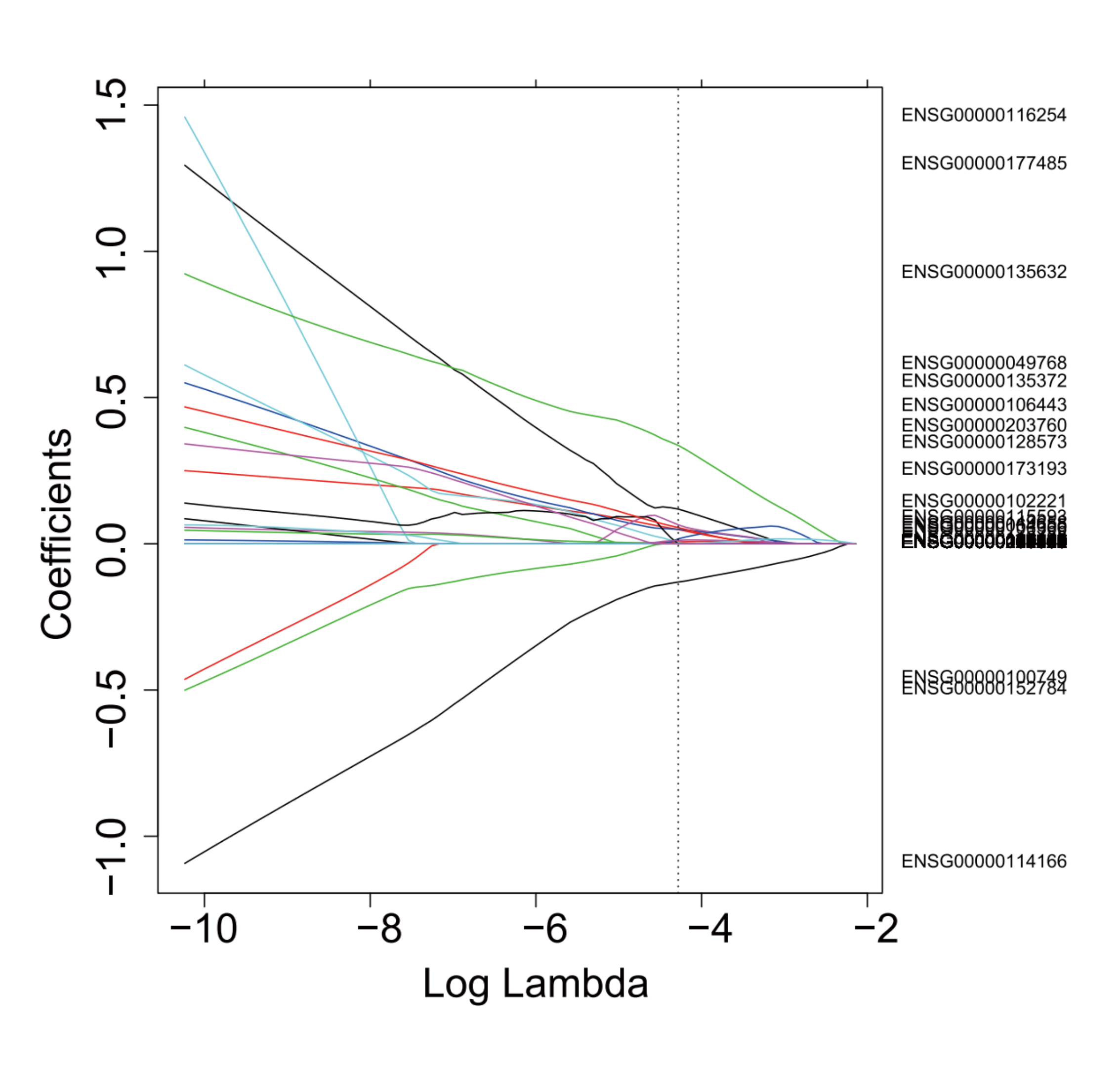
Candidate markers identified by the selection model
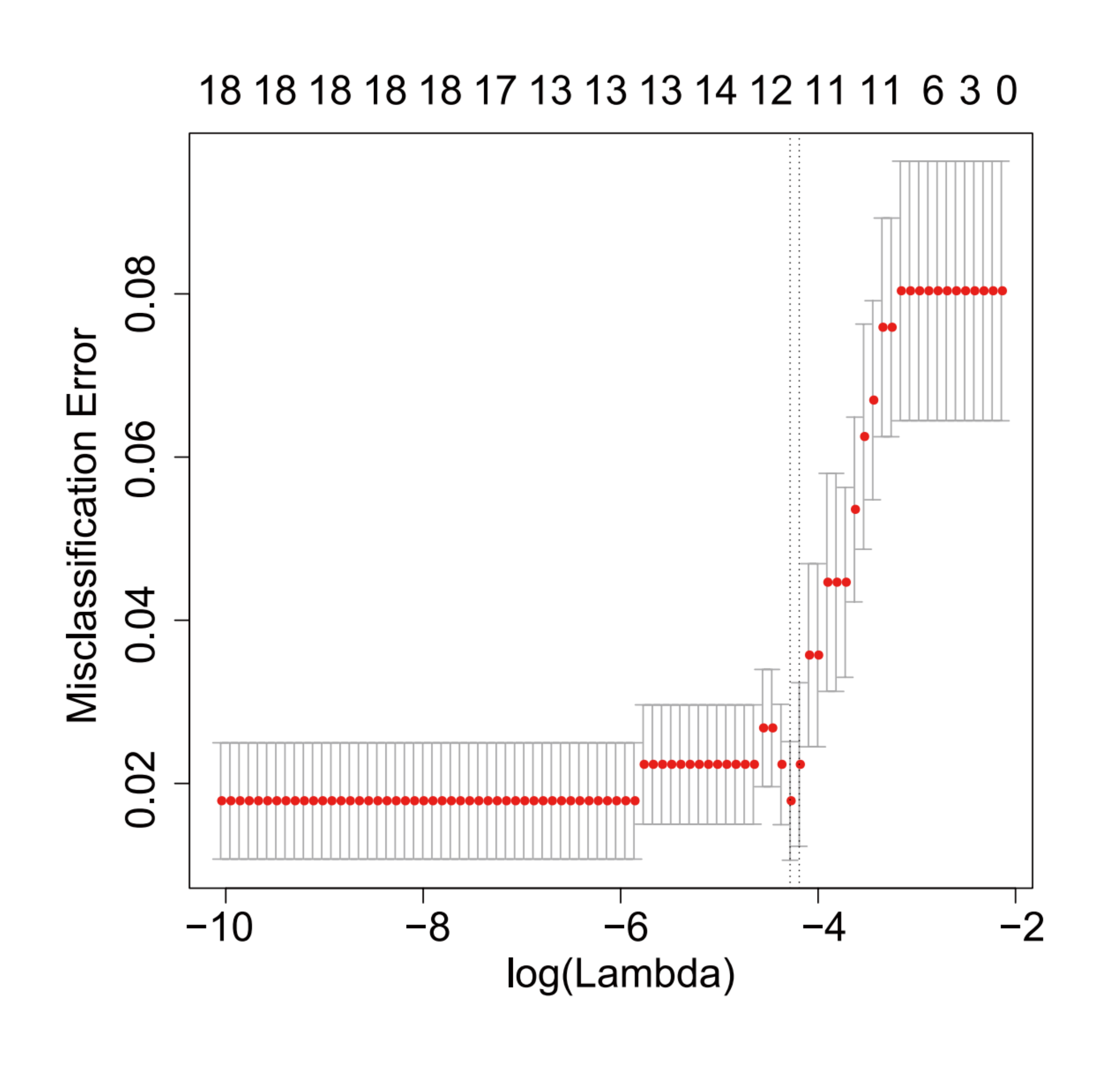
Optimal hyperparameters for the selection model
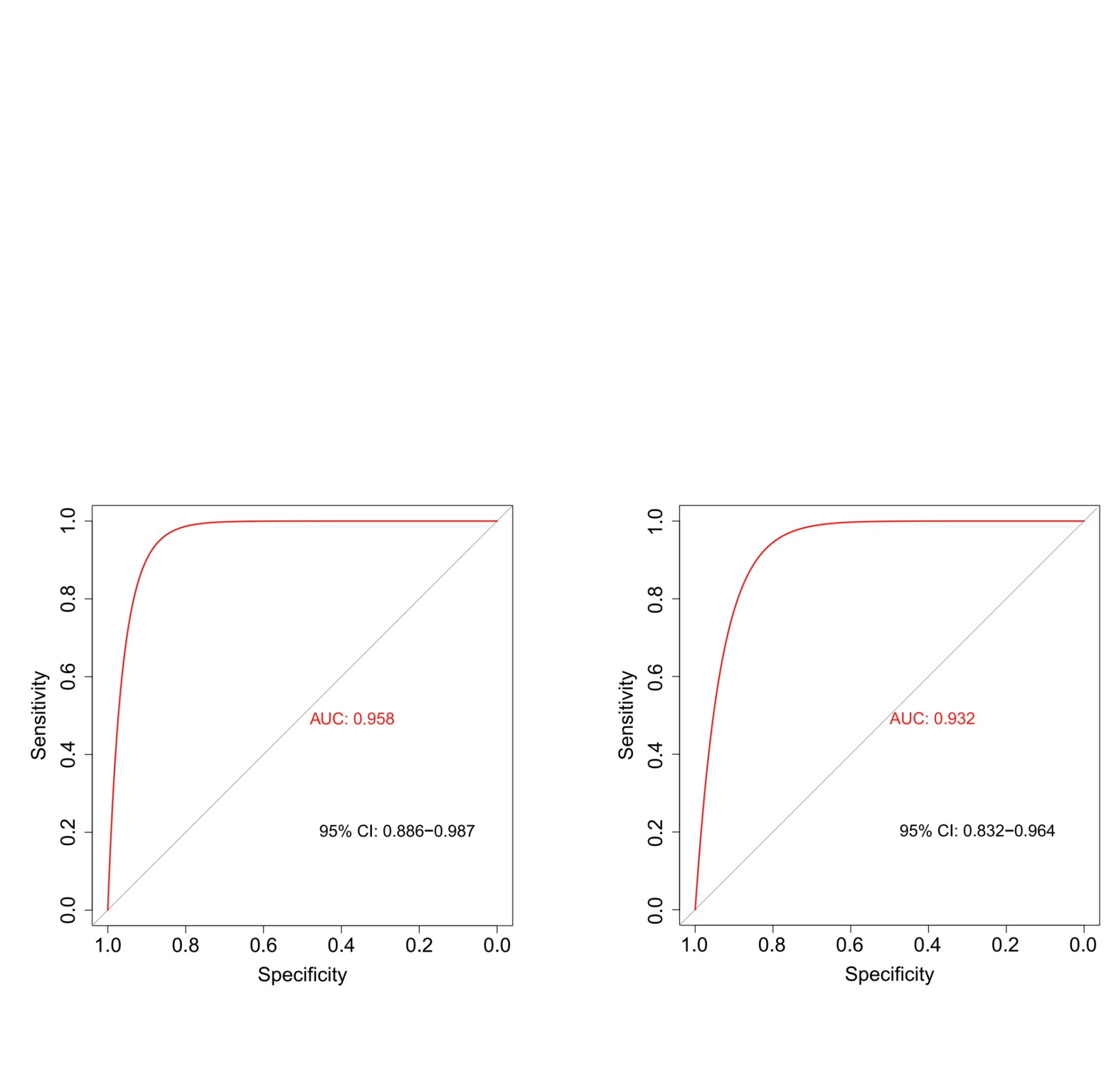
ROC Analysis
