PAR-CLIP-seq
PAR-CLIP-seq
PAR-CLIP-seq is an advanced technique based on the incorporation of photoreactive ribonucleoside analogs for crosslinking and immunoprecipitation (1). By incorporating 4-thiouridine (4sU) into nascent RNA and analyzing specific Thymine-to-Cytosine (T>C) transition sites during sequencing, this method enables the genome-wide investigation of RNA-protein interactions and the precise identification of RNA Binding Protein (RBP) binding sites.
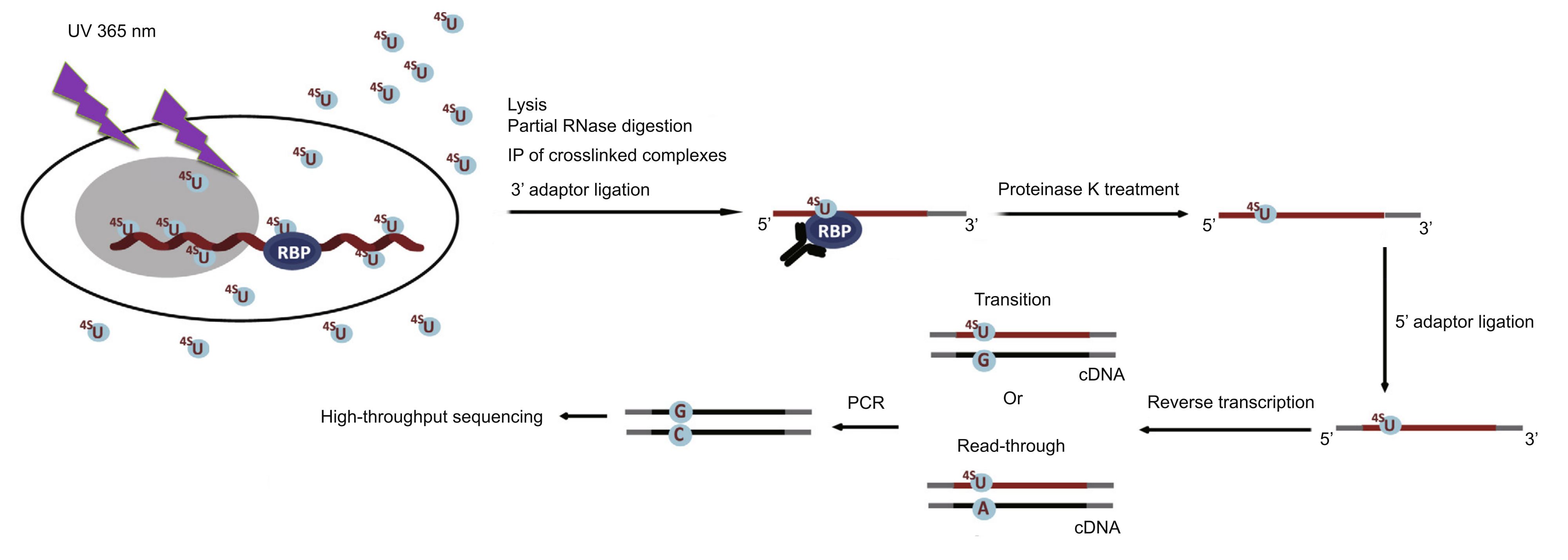
Technical Principles
The process involves incorporating photoreactive nucleoside analogs, such as 4sU, into the nascent RNA transcripts of live cells. Upon irradiation with 365 nm UV light, the 4sU-labeled RNA is induced to crosslink specifically with interacting RBPs. The target RBP is then immunoprecipitated, and the crosslinked RNA is isolated.
The RNA is subsequently converted into a cDNA library for deep sequencing. The 4sU treatment causes characteristic T-to-C transitions during reverse transcription at the crosslinking sites. Consequently, PAR-CLIP-seq allows for the accurate localization of the specific protein-RNA interaction interface by mapping these mutation signatures.
Advantages
1. High Crosslinking Efficiency: 100–1,000 times higher efficiency compared to standard CLIP-seq methods.
2. High Resolution: Capable of pinpointing the exact binding sites.
3. High Specificity: Reduced background noise through specific photoactivation.
Applications
1. Binding Site Identification: Identification of direct binding sites between proteins and RNA across the entire transcriptome.
2. Interaction Mapping: Elucidation of genome-wide RNA-protein interaction networks.
Sample Requirements
Type
Live cells
Input
≥3×10⁷ cells/sample
Bioinformatics Analysis
1. Data Processing: Raw data quality control (QC) and filtering.
2. Alignment: Mapping reads to the reference genome.
3. Peak Calling: Identification of enriched binding regions.
4. Annotation: Genomic annotation of identified peaks.
5. Functional Analysis: GO (Gene Ontology) enrichment analysis.
6. Pathway Analysis: KEGG pathway enrichment analysis.
7. Motif Analysis: Identification of consensus binding sequences.
8. Signal Matrix Analysis: Generation of peak heatmaps.
9. CIMS Analysis: Crosslinking Induced Mutation Sites analysis to pinpoint binding at single-nucleotide resolution.
10. Visualization: Generation of track files (e.g., for IGV).
11. Differential Analysis: Identification of differential binding sites (requires multi-group experimental design).
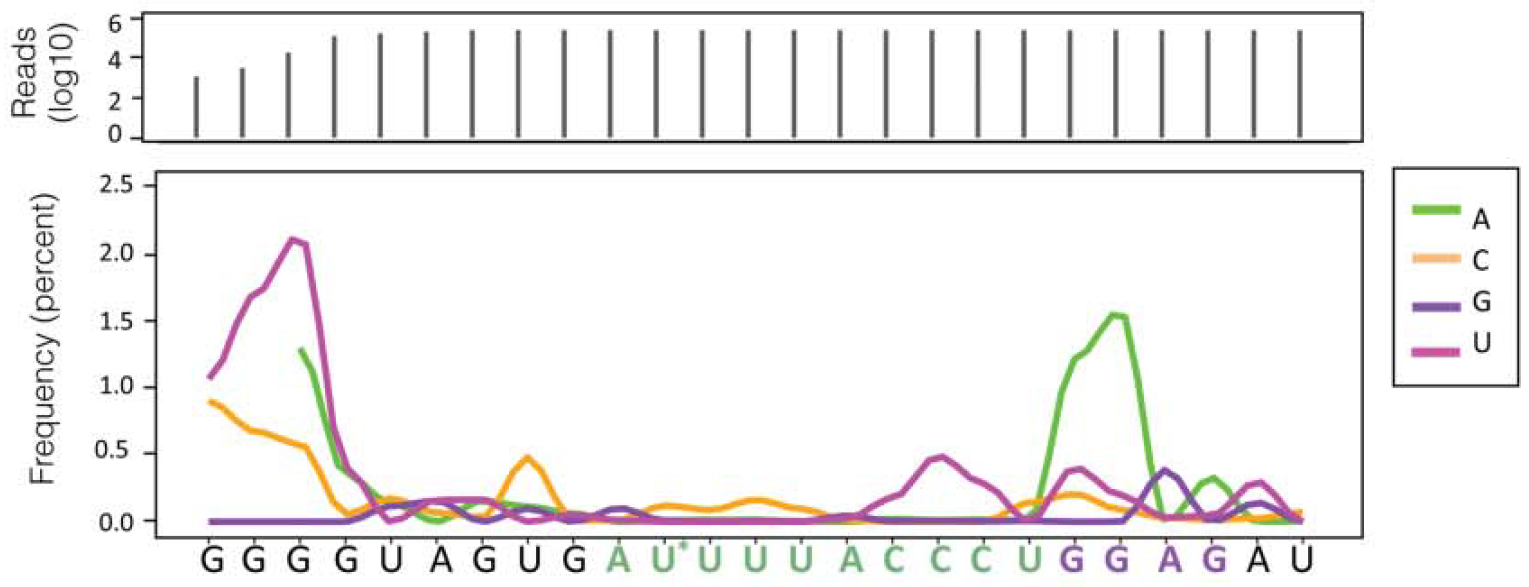
CIMS Analysis showing mutation frequency
Demo
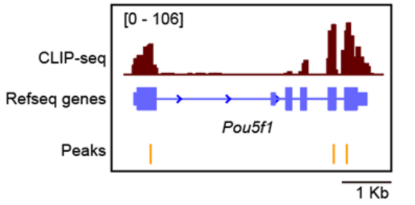
IGV Visualization of Peaks

Motif Analysis
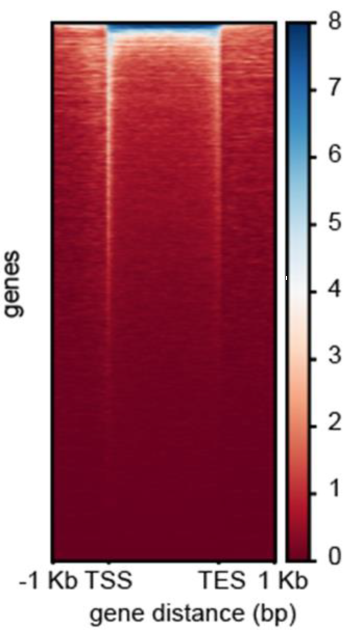
Signal Distribution Heatmap
