RADICL-seq: RNA-DNA interactions
Introduction
RNA and DNA Interacting Complexes Ligated and Sequenced (RADICL-seq) [5] is a novel technology for exploring RNA-chromatin interactions, map the interaction landscape between RNA and chromatin within the nucleus, identify genome coverage patterns across different transcript classes, and detect cell-specific RNA-chromatin interactions. RADICL-seq allows for the capture of RNA that interacts with genomic target regions and provides a comprehensive localization for each captured RNA molecule.
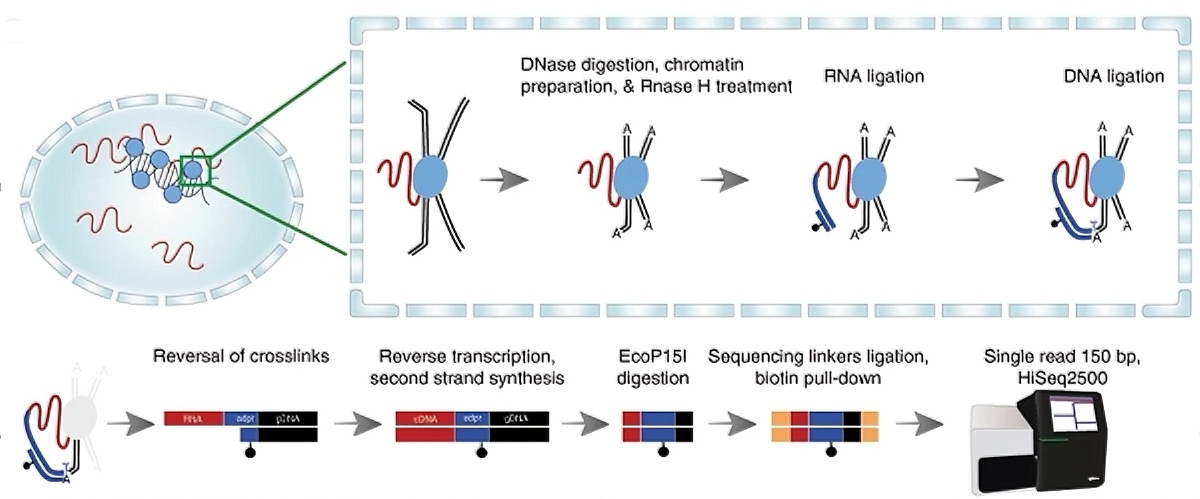
Figure 1. Workflow of RADICL-seq (Bonetti A, et al. Nat Commun. 2020)
Technical principle
The RADICL-seq workflow begins by fixing RNA, proteins, and DNA in situ within the nucleus with formaldehyde. A specifically designed linker is then ligated to juxtaposed RNA and DNA molecules. After reversal of cross-links and protein removal, only the linker-associated RNA–DNA hybrids are recovered and converted into double-stranded DNA for next-generation sequencing. Reads are finally aligned to the reference genome to map RNA–DNA interactions.
Key steps of the RADICL-seq protocol:
1. Cross-linking: Living cells are treated with 1 % formaldehyde to covalently stabilize endogenous RNA–protein–DNA complexes.
2. Nuclei isolation: Intact nuclei are purified to enrich for chromatin-associated RNA.
3. Partial DNase I digestion: Genomic DNA is fragmented under controlled conditions to generate accessible ends adjacent to RNA contact sites.
4. rRNA depletion: Ribosomal RNA is selectively digested with RNase H to increase the fraction of informative reads.
5. RNA–DNA linker ligation: A bifunctional adaptor is ligated to the juxtaposed 3′ end of RNA and the 5′ end of DNA.
6. Reversal of cross-links and protein removal: Formaldehyde cross-links are reversed, proteins are degraded, and RNA–DNA chimeras are purified.
7. Library construction and sequencing: RNA–DNA hybrids are reverse-transcribed and converted into double-stranded DNA libraries, which are then sequenced on a next-generation platform
Bioinformatic Analysis
Applications
1.Systematic mapping of genome-wide RNA–DNA interactions within the nucleus.
2.Elucidation of RNA-mediated transcriptional regulatory mechanisms.
3.Construction of three-dimensional genome regulatory networks.
4.Decoding dysregulation caused by non-coding enhancer mutations.
Analysis Content
1. Quality control of sequencing data
2. Linker alignment
3. RNA/DNA separation and data quality control
4. Genomic alignment
5. RNA-DNA interaction matrix
6. Genome-wide RNA-DNA interaction mapping
7. RNA-DNA interaction enhancer network (Advanced analysis)
Sample Requirements
Formaldehyde crosslinked cells, ≥ 2×10⁶ cells/sample
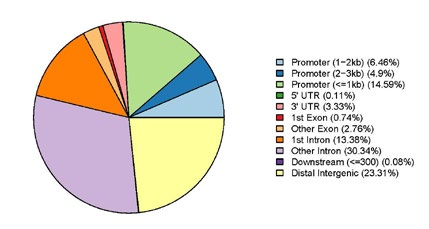
Figure 2. Interaction type diagram
Demo

Figure 3. RNA-DNA circular interaction diagram
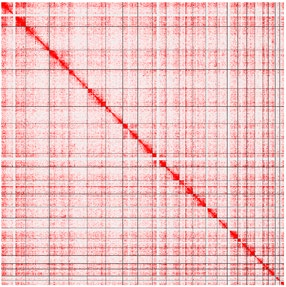
Figure 4. RNA-DNA global interaction map
Related Articles
Nat Commun: RADICL-seq identifies general and cell type–specific principles of genome-wide RNA-chromatin interactions (1)
RADICL-seq leverages proximity-based cross-linking and ligation to markedly reduce the bias toward nascent transcripts that is inherent in existing approaches, while simultaneously increasing genomic coverage and unique mapping efficiency. The method can delineate transcript class–specific “occupancy” patterns across the genome, identify cell-type-specific RNA–chromatin interactions, and illuminate the contribution of transcription to higher-order chromatin architecture. The resulting datasets provide a comprehensive view of how chromatin is organized and regulated, and enable a global assessment of lncRNA function.
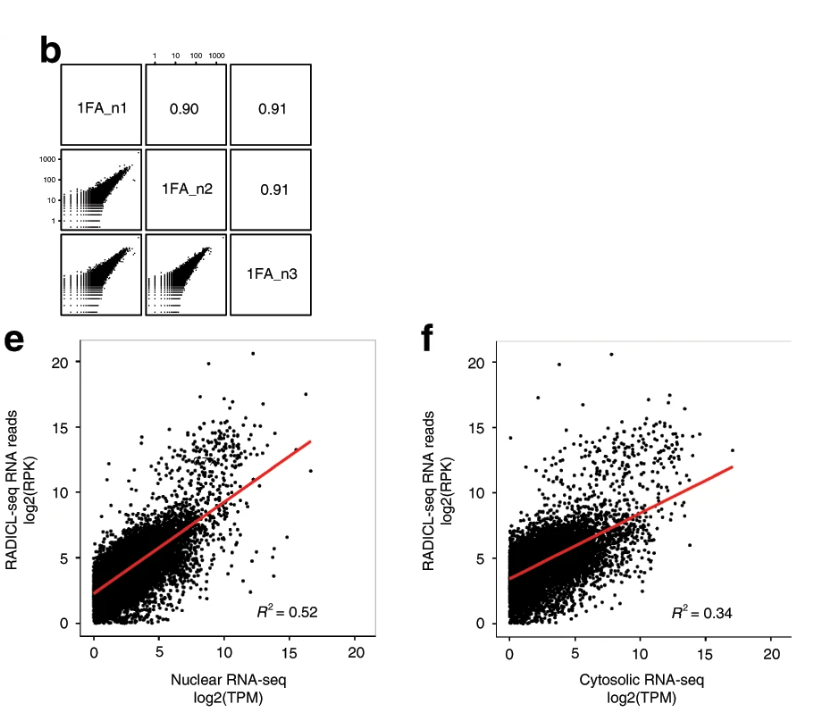
1. Identify RNA-DNA interactions in the nucleus and capture more nuclear information.
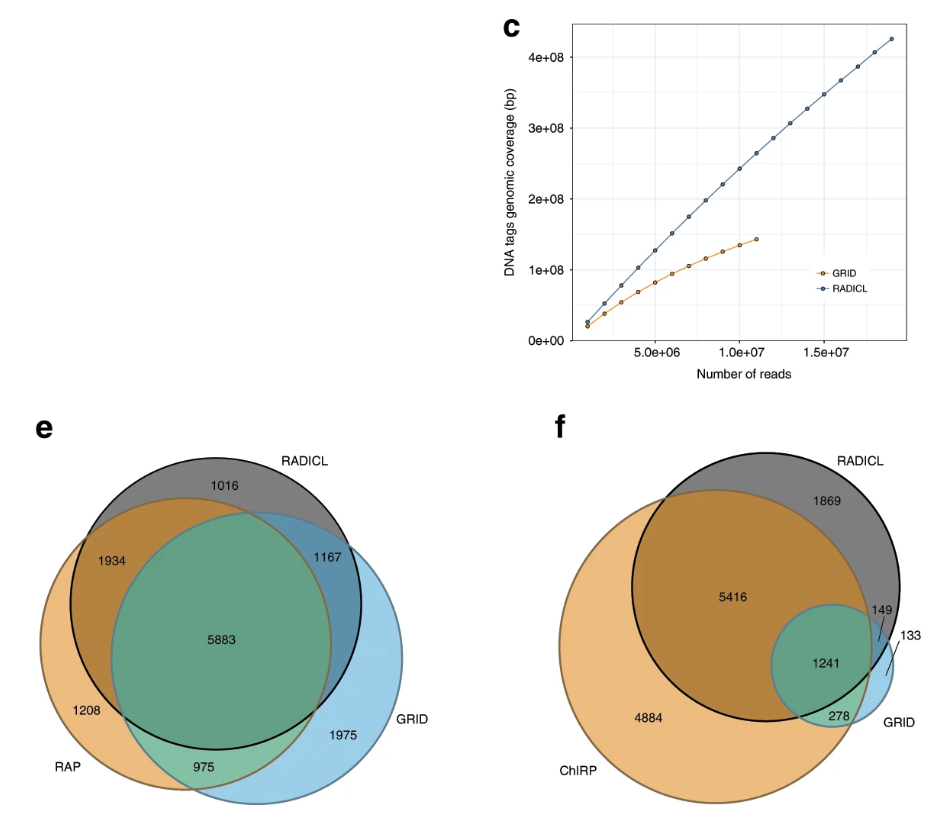
2. RADICL-seq has higher resolution, lower sample size, and higher capture rate.
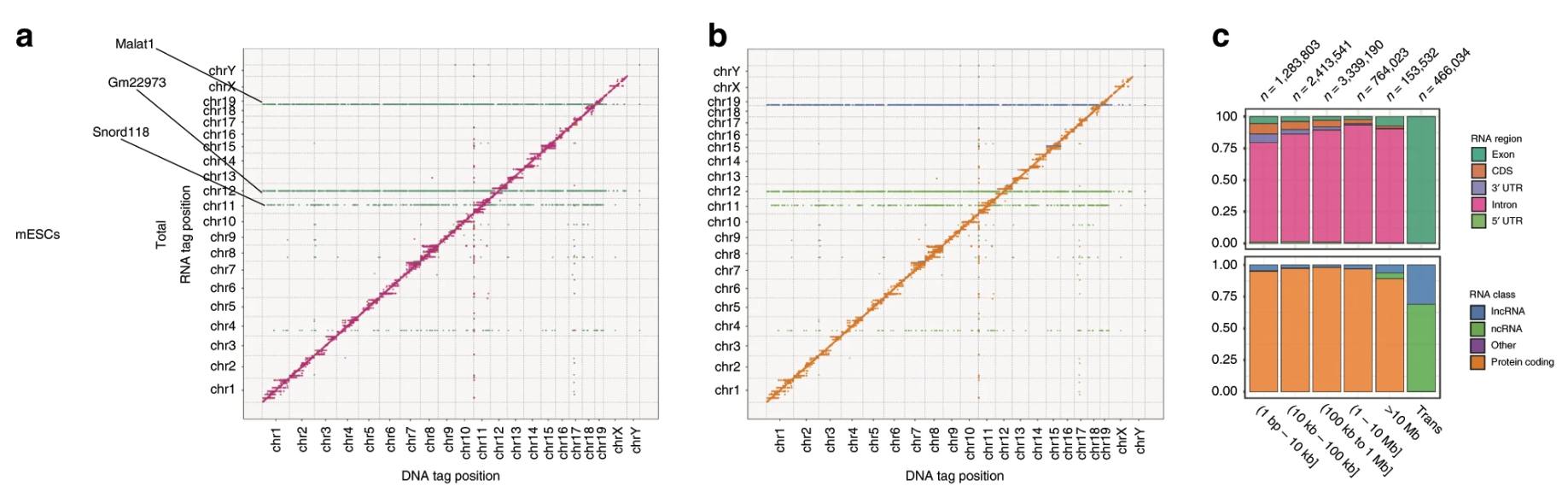
3. Identification of genome-wide RNA-DNA interactions with rich data comparable to Hi-C
