Epi-SELECT™ m6A Detection Kit
Epi-SELECT™ m6A Detection Kit
single-base elongation-and ligation-based qPCR amplification method
The Single - base Elongation - and Ligation - based qPCR amplification method (SELECT) technology was developed by the research team led by Professor Guifang Jia at Peking University. This innovative approach enables the detection of m6A modifications with single - base resolution in low - abundance transcripts within a mere three hours. In contrast to conventional methods, SELECT permits the targeted quantification of m6A at specific sites without the necessity of antibody enrichment. It facilitates the analysis of disparities in modification abundance among samples and validates the presence of m6A at target loci.
EPIBIOTEK® has obtained an exclusive global licensing agreement from Peking University for the SELECT technology. We provide a comprehensive set of SELECT detection kits, which addresses the market demand for routine, site - specific m6A detection. This tool aids researchers in identifying the targets of m6A - related proteins and clarifying the underlying mechanisms of m6A modifications, thereby laying the foundation for novel discoveries in RNA m6A biomarkers.
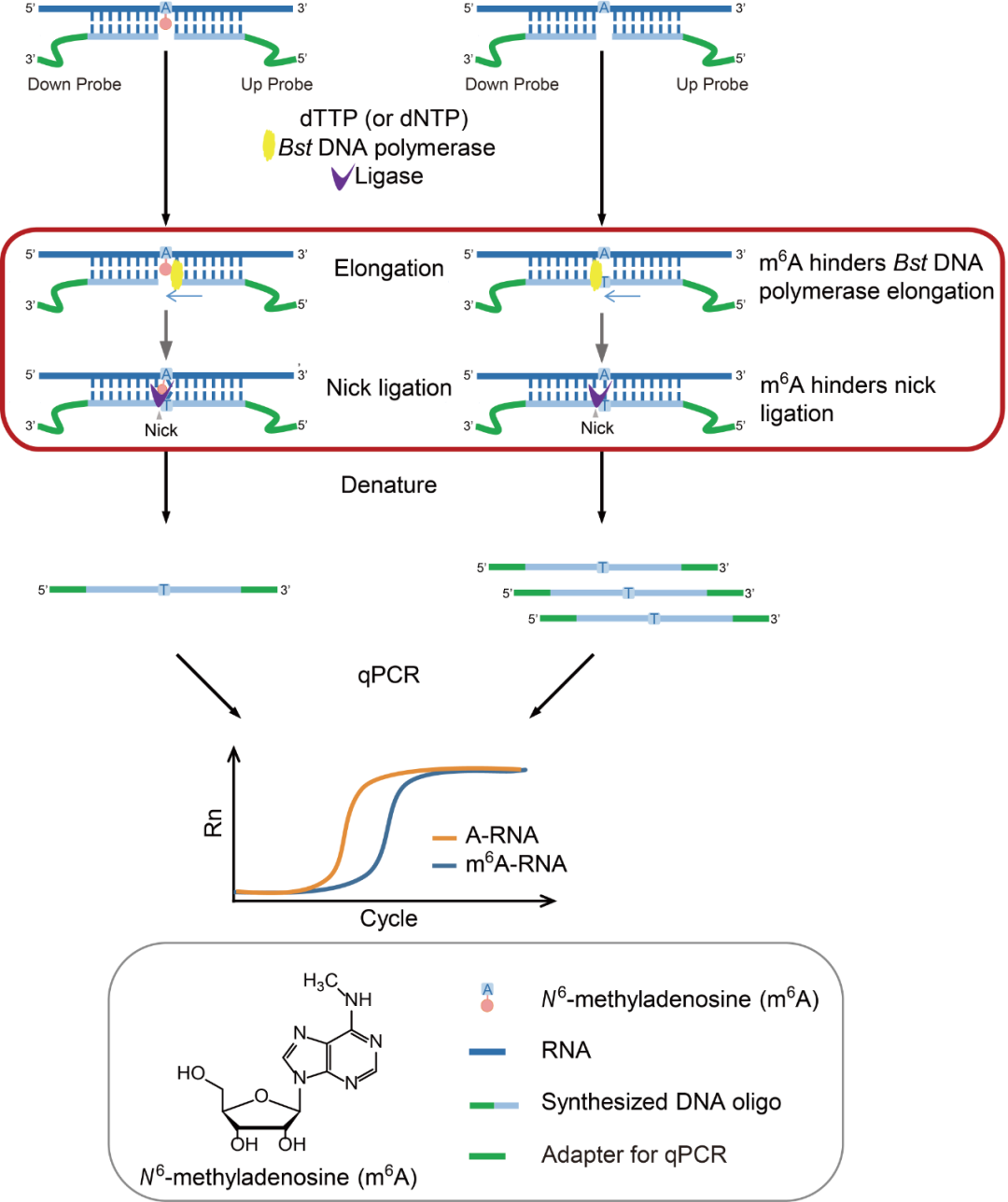
Workflow
Applications
1. Differential Analysis: Verify differences in m6A modification abundance at specific sites between samples.
2. Site Validation: Detect the presence of m6A modifications at specific RNA target sites.
3. Absolute Quantification: Determine the absolute fraction of m6A modification at specific sites.
4. Target Identification: Identify functional targets of m6A-associated proteins (methyltransferases, demethylases, readers).
Advantages
1. High Precision: Utilizes fluorescent qPCR to achieve single-base resolution.
2. Cost-Effective & Accessible: Antibody-free and isotope-free workflow significantly reduces costs and simplifies operation.
3. High Sensitivity: Requires as little as 1 μg of total RNA input.
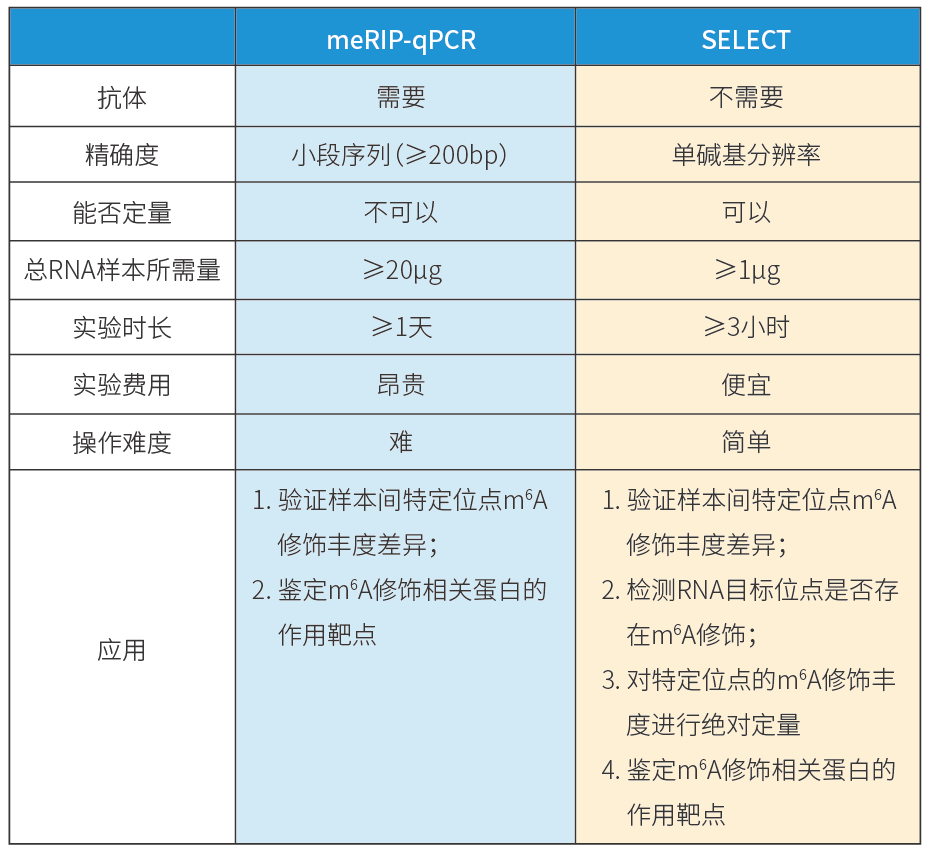
Technical Comparison
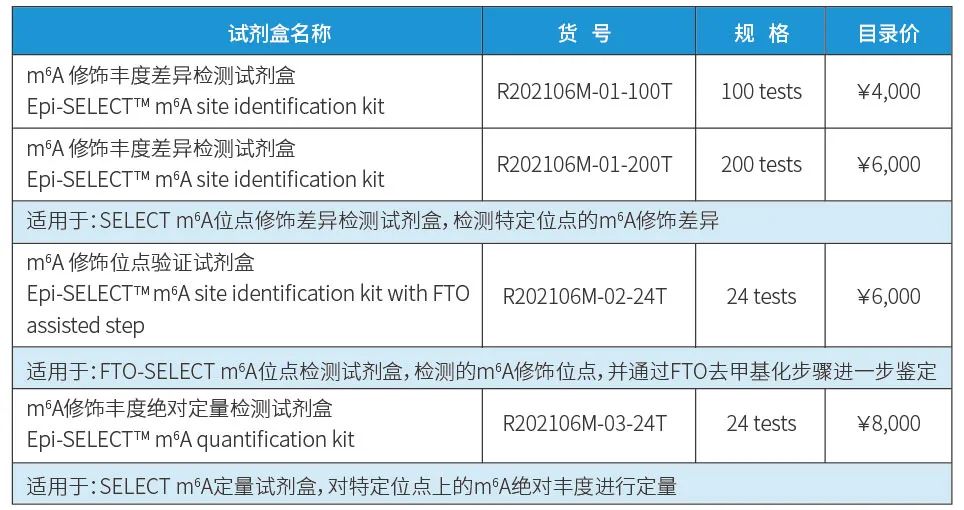
Kit Information
EPIBIOTEK® Publications
[1] Yu L, Alariqi M, Li B, et al. CRISPR/dCas13(Rx) Derived RNA N6-methyladenosine (m6A) Dynamic Modification in Plant. Adv Sci (Weinh). 2024;11(39):e2401118.
[2] Yu H, Li QS, Guo JN, et al. METTL14-mediated m6A methylation of pri-miR-5099 to facilitate cardiomyocyte pyroptosis in myocardial infarction. Acta Pharmacol Sin. Published online February 12, 2025.
[3] Chen, H., Zhao, X., Yang, W. et al. RNA N6-methyladenosine modification-based biomarkers for absorbed ionizing radiation dose estimation. Nat Commun 14, 6912 (2023).
[4] Huang J, Jiang B, Li GW, et al. m6A-modified lincRNA Dubr is required for neuronal development by stabilizing YTHDF1/3 and facilitating mRNA translation. Cell Rep. 2022 Nov 22;41(8):111693.
[5] Zhang S, Lv C, Niu Y, et al. RBM3 suppresses stemness remodeling of prostate cancer in bone microenvironment by modulating N6-methyladenosine on CTNNB1 mRNA. Cell Death Dis. 2023 Feb 7;14(2):91.
[6] Tan L, Qin Y, Xie R, et al. N6-methyladenosine-associated prognostic pseudogenes contribute to predicting immunotherapy benefits and therapeutic agents in head and neck squamous cell carcinoma. Theranostics 2022;12(17).
[7] Chen X, Wang Y, Wang JN, et al. m6A modification of circSPECC1 suppresses RPE oxidative damage and maintains retinal homeostasis. Cell Rep. 2022 Nov 15;41(7):111671.
[8] Sheng R, Meng W, Zhang Z, et al. METTL3 regulates cartilage development and homeostasis by affecting Lats1 mRNA stability in an m6A-YTHDF2-dependent manner. Cell Rep. 2024;43(8):114535.
[9] Zhou J, Tang J, Zhang C, et al. ALKBH5 targets ACSL4 mRNA stability to modulate ferroptosis in hyperbilirubinemia-induced brain damage. Free Radic Biol Med. 2024;220:271-287.
[10] Cai B, Ma M, Yuan R, et al. MYH1G-AS is a chromatin-associated lncRNA that regulates skeletal muscle development in chicken. Cell Mol Biol Lett. 2024 Jan 4;29(1):9.
[11] Li L, Cheng H, Zhou Y, et al. METTL3 regulates autophagy of hypoxia-induced cardiomyocytes by targeting ATG7. Cell Death Discov. 2025;11(1):37. Published 2025 Feb 1.
[12] Liu K, Wang X, Wang J, et al. N 6-methyladenosine modifications stabilize phosphate starvation response-related mRNAs in plant adaptation to nutrient-deficient stress. Nat Commun. 2025;16(1):4093. Published 2025 May 1.
