Polysome-seq
Polysome-seq
Polysome profiling
The translatome serves as the critical functional link between the transcriptome and the proteome. Polysome profiling is the classic technique (1) and the recognized "gold standard" for assessing translation efficiency by utilizing the high sedimentation coefficient of ribosomes to separate polysomes.
Polysome-seq employs the Biocamp automated density gradient fractionation system. This service separates polysomes via sucrose density gradients and generates precise polysome profiles. Combined with high-throughput sequencing, it identifies genes with differential Translational Efficiency (TE), facilitating in-depth research into regulatory mechanisms occurring between mRNA transcription and protein synthesis.
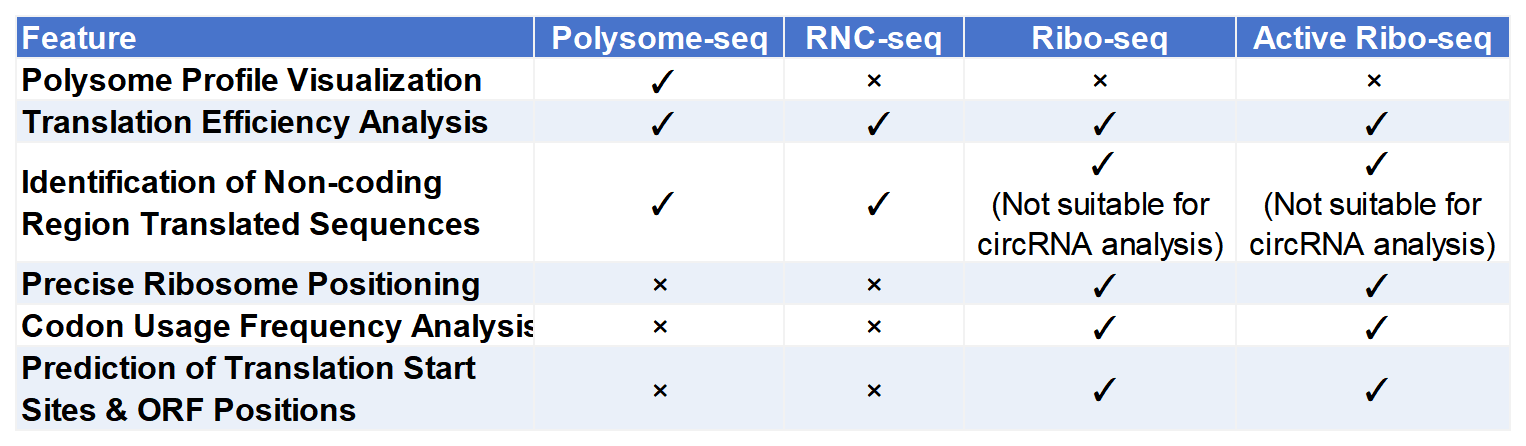
Comparison of Translatome Technologies
Applications (Combined with longRNA-seq)
1. Determine protein translation efficiency.
2. Investigate translational regulation and gene expression, such as target mRNA degradation and translation inhibition.
3. Explore the regulatory mechanisms of epigenetic modifications.
4. Identify co-translational degradation of nascent peptides.
5. Study translation-related events, such as ribosome synthesis, assembly, and translation-associated factors.
Advantages
1.Gold standard for translation efficiency via polysome isolation.
2.Direct visualization of cellular translation status.
3.Quantifies ribosome load per mRNA.
4.Detects non-coding region translation and regulation.
5.Compatible with diverse sample types.
Sample Requirements
Validated for Human, Mouse, and Rat. (Please consult for low-input samples or other species assessments)
1. Grouping: A minimum of 2 groups is required (e.g., Control vs. Experimental; Normal vs. Patient).
2. Pairing: Both Translatome (Polysome-bound RNA) and Transcriptome (Total RNA) sequencing must be performed for each sample to calculate TE.
3. Replicates: Recommended biological replicates: 3 vs. 3.
Cells
≥1×10^7 cells/sample
Tissue
≥100 mg/sample
Bioinformatics Analysis
Polysome Profiling Analysis
1.Raw data filtering and sequencing QC.
2.Removal of rRNA/tRNA and read length filtering.
3.Alignment to Reference Genome and Transcriptome.
4.Polysome Profile Generation: Visualization of the gradient trace.
5.Global Translational Activity Analysis: Calculation of the Polysome-to-Monosome (P/M) ratio.
6.Differential Translational Gene Analysis.
7.Heatmap of Differentially Translated Genes (Requires biological replicates).
8.GO and KEGG Enrichment Analysis for Differentially Translated Genes.
9.TE Calculation: Requires matched mRNA-seq or longRNA-seq data.
10.Differential TE Analysis.
11.GO and KEGG Enrichment Analysis for Differential TE Genes.
Control RNA-seq / longRNA-seq Analysis
1.Adapter removal and QC.
2.Reference Genome Alignment.
3.Gene Expression Quantification.
4.Gene Annotation.
5.Differential Expression Analysis.
6.Heatmap of Differentially Expressed Genes (Requires biological replicates).
7.GO and KEGG Enrichment Analysis.
