Ribosome Profiling (Ribo-seq)
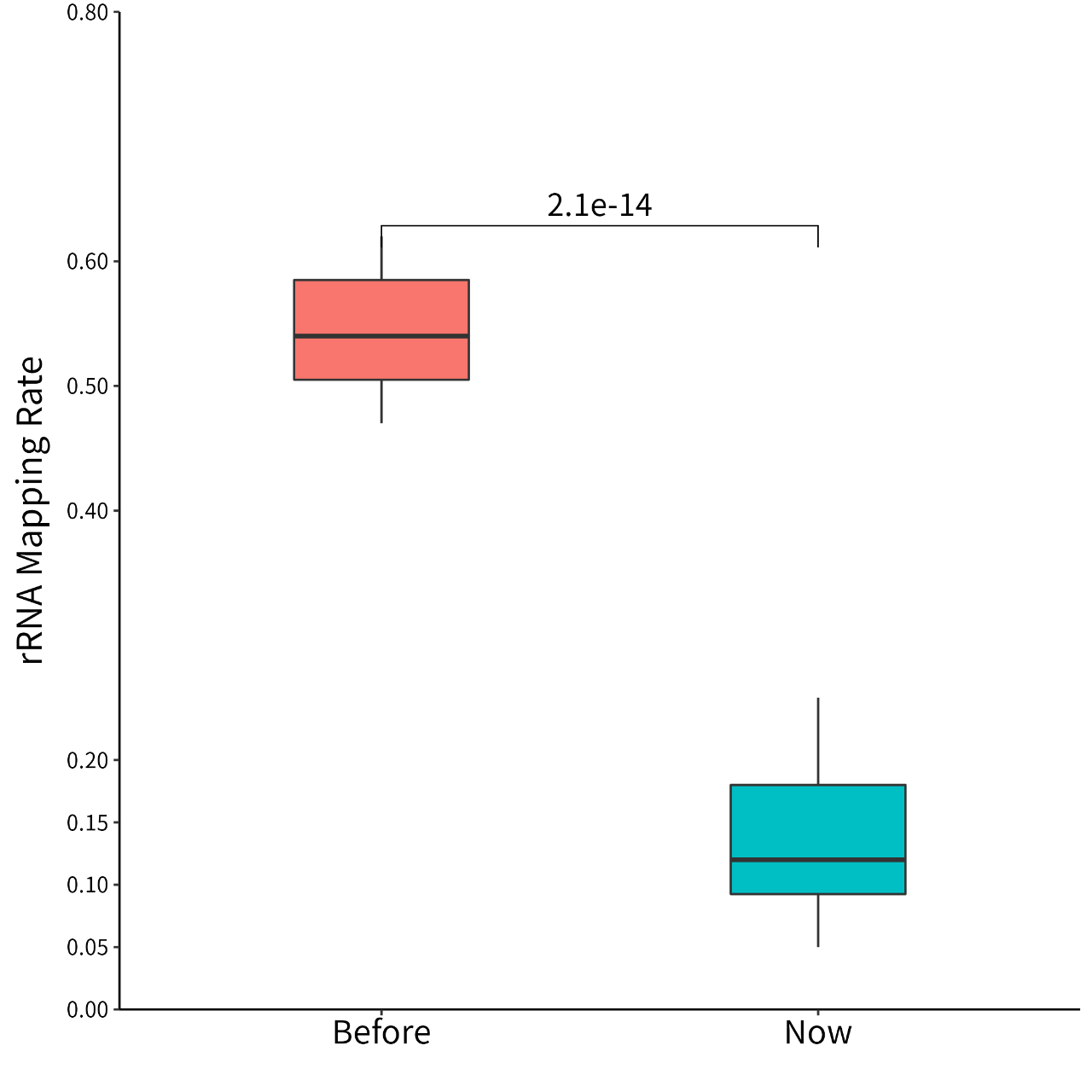
Figure 1. For Ribo-seq experimental data, the average proportion of rRNA was reduced from 55% to 14%.
Introduction
Ribo-seq
Ribosome Profiling (Ribo-seq) is a translational genomics research technology first published by the Weissman laboratory in 2009 (1). Researchers can use Ribo-seq to detect the translation status of proteins at the genome level and obtain comprehensive, high-quality information on protein translation rates. It helps in understanding protein expression and abundance and allows for the direct study of the translation process.
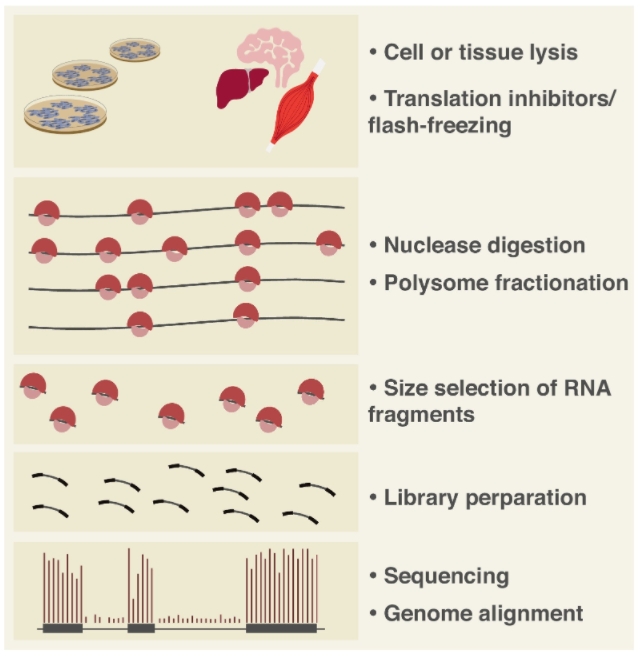
Figure 2. Workflow of Ribo-seq (2)
Ribosome-nuclease treatment degrades any mRNA regions not protected by translating ribosomes. High-throughput sequencing of the surviving fragments—termed ribosome footprints—maps the exact positions of ribosomes along the transcript. Regions with higher footprint density correspond to sites of slower elongation kinetics.
Applications
When integrated with longRNA-seq, Ribo-seq enables comprehensive interrogation of active translation on individual transcripts, providing unprecedented insight into the most fundamental layer of gene regulation:
1. Profiling ribosome occupancy and translational activity across transcripts.
2. Inferring translation start sites and open reading frame (ORF) locations.
3. Quantifying protein translation efficiency.
4. Investigating the interplay between translational control and overall gene expression.
5. Discovering novel proteins and short peptides.
Sample Requirements
Limited to human, mouse, and rat cells and tissues; other species require evaluation.
Types & Amount
1. Cells, ≥ 51×10⁷ cells/sample
2. Tissues, ≥ 50 mg/sample
Sample Grouping
1. At least 2 groups of samples are required, including a control group and an experimental group (clinical samples are normal group and patient group)
2. Each sample was subjected to Ribo-seq and longRNA-seq
3. Sample size suggestion: 3 VS 3
Bioinformatic Analysis
Basic Analysis (Ribo-seq )
1. Raw data filtering and sequencing quality assessment
2. Removal of rRNA and tRNA, and filtering of read length
3. Alignment to the reference genome and reference transcriptome
4. Ribosome profiling quality control (read length distribution, RFs distribution, 3nt periodicity feature)
5. Genome-wide translational activity analysis
6. Differential translation gene analysis
7. Heatmap of differential translation efficiency (only with biological replicates)
8. GO analysis of differentially translated genes
9 KEGG analysis of differentially translated genes
10. Calculation of gene translation efficiency (requires mRNA-seq or longRNA-seq)
11. Analysis of differential translation efficiency
12. GO analysis of genes with differential translation efficiency
13. KEGG analysis of genes with differential translation efficiency
14. Codon usage frequency analysis
15. ORF prediction
16. Translation pause analysis
Control Analysis (long RNA-seq )
1. Raw reads trimming and quality control
2. Alignment to the reference genome
3. Quantitative analysis of gene expression
4. Gene annotation
5. Differential expression analysis
6. Heatmap of differentially expressed genes (only with biological replicates)
7. GO analysis of differentially expressed genes
8. KEGG analysis of differentially expressed genes
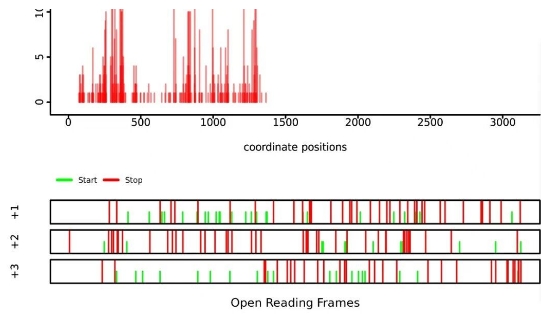
Figure 3. Translation pause analysis
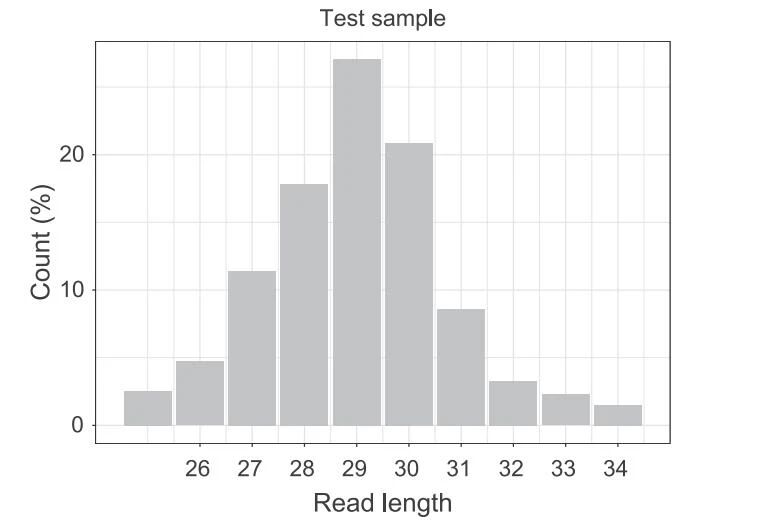
Figure 4. Length distribution of Ribo-seq reads aligned to the genome
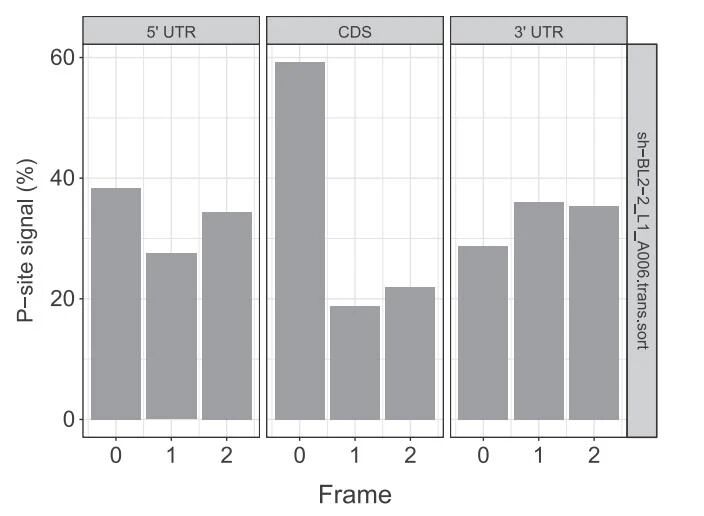
Figure 5. Distribution of P-site signals in 5 'UTR, CDS, and 3' UTR
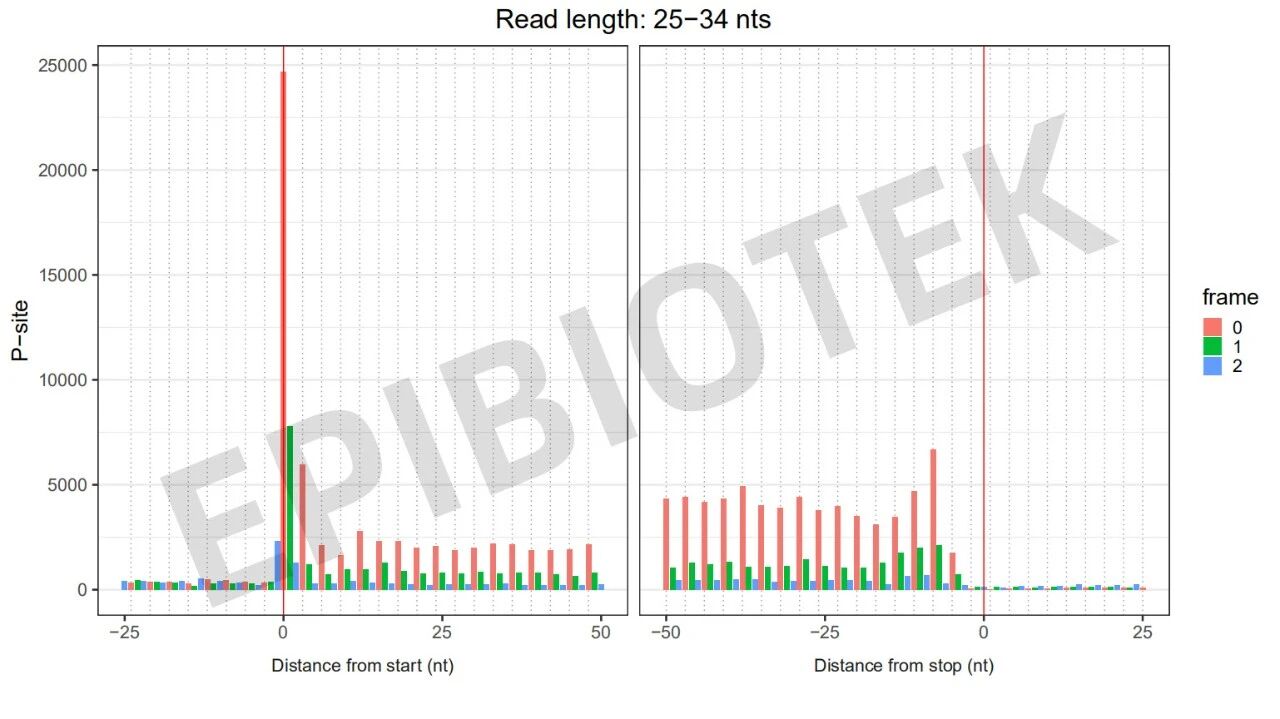
Figure 6. 3nt quality control diagram of cell precipitation samples
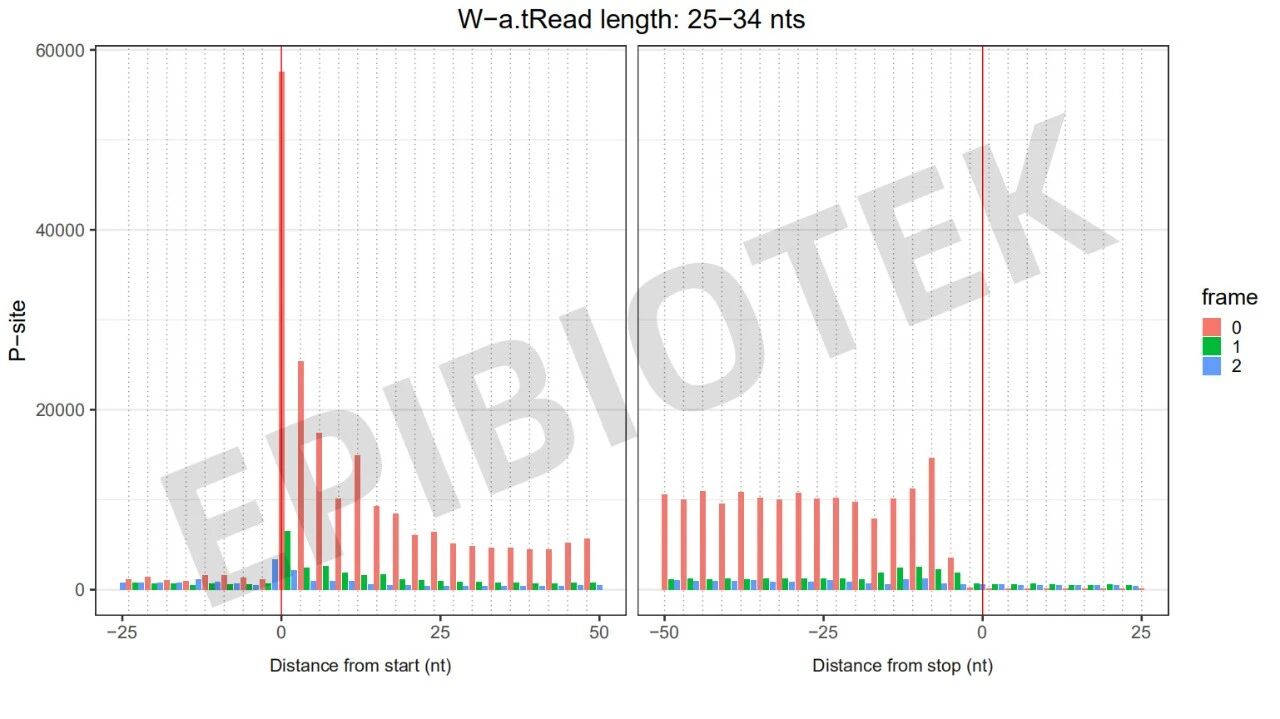
Figure 7. 3nt quality control diagram of live cell samples on the wall
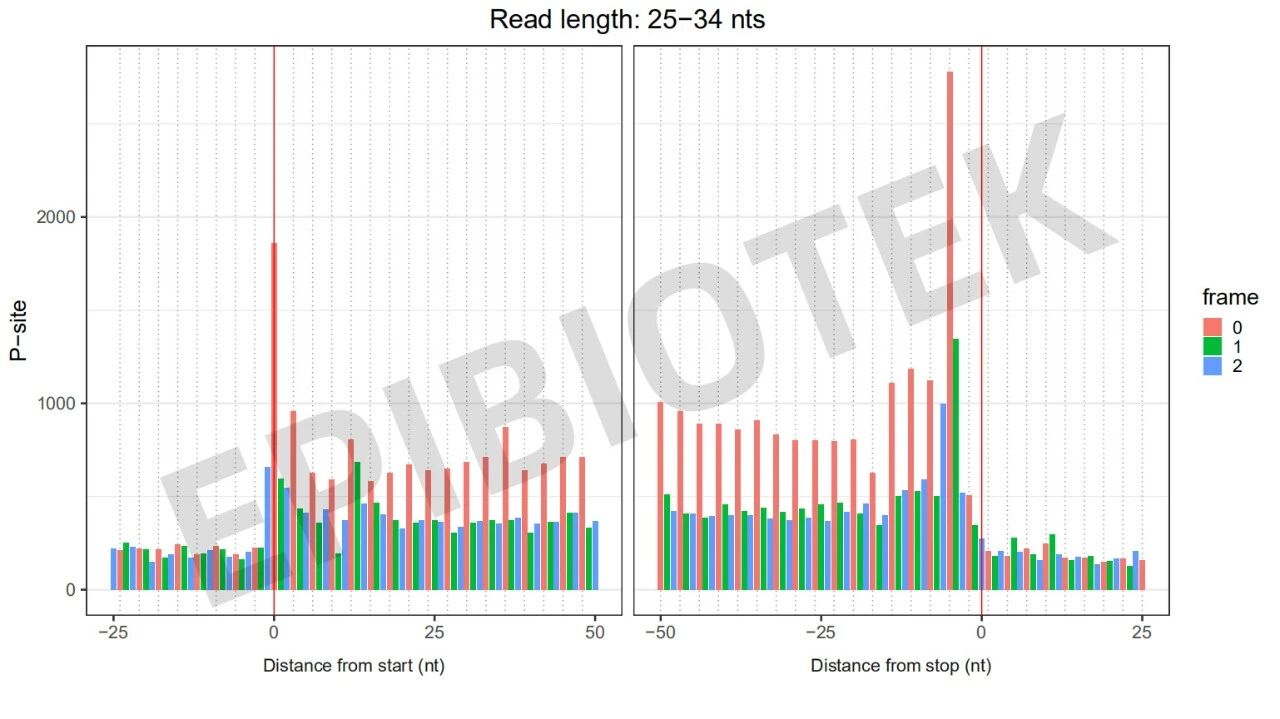
Figure 8. 3nt quality control diagram of tissue samples
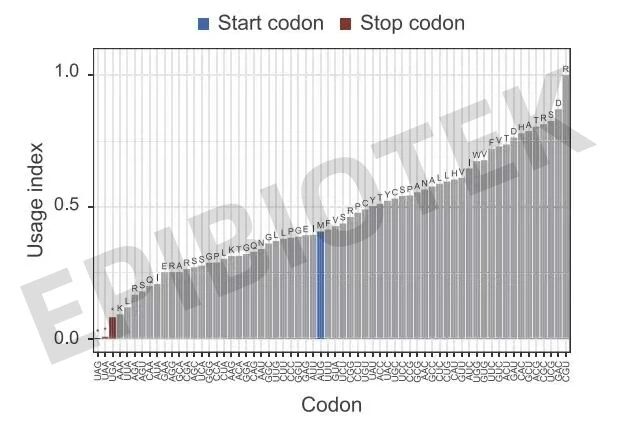
Figure 9. Analysis of different codon usage frequencies
EPIBIOTEK® Project Publications
1. Liu J, Yan X, Wu H, et al. RNA codon expansion via programmable pseudouridine editing and decoding. Nature. 2025;643(8074):1410-1420.
2. Gu J, Li Y, Chen Q, et al. Tumor-produced ammonia is metabolized by regulatory T cells to further impede anti-tumor immunity. Cell. 2026;189(2):418-434.e24.
3. Rao H, Wang A, Xu Y, et al. Ribosome Homeostasis Regulated by SETD2 Preserves Intestinal Epithelial Barrier. Adv Sci (Weinh). Published online January 4, 2026.
4. Shi L, Zhang M, Yang H, et al. NAT10 regulates heart development and function by maintaining the expression of genes related to fatty acid β-oxidation and heart contraction. Cell Death Differ. Published online September 13, 2025.
5. Li K, Guo C, Wu X, et al. Enhanced Protein Synthesis and Hippocampus-Dependent Memory via Inhibition of YTHDF2-Mediated m6A mRNA Degradation. Adv Sci (Weinh).
6. Zhang XY, Zhang YH, Liang NN, et al. The m6A modification-mediated upregulation of ETS2 translation drives arsenic-induced spermatogonial senescence. Free Radic Biol Med. 2025;240:384-396.
7. Xiao Z, Wei X, Li P, et al. Impact of N-acetyltransferase 10 on macrophage activation and inflammation-induced cardiac dysfunction. Cell Death Dis. 2025;16(1):471. Published 2025 Jul 1.
8. Liu WC, Wei YH, Chen JF, et al. Inhibition of tumor-intrinsic NAT10 enhances antitumor immunity by triggering type I interferon response via MYC/CDK2/DNMT1 pathway. Nat Commun. 2025;16(1):5154. Published 2025 Jun 3.
9. Niu FW, Liu MD, Yao K, et al. Mitochondrial ROS-associated integrated stress response is involved in arsenic-induced blood-testis barrier disruption and protective effect of melatonin. Environ Int. 2025;197:109346.
10. Shu X, Wang R, Liu Y, et al. S6K1 overexpression enhances autophagy in breast cancer cells by inducing the translation of CLU. Chin Med J (Engl). Published online May 23, 2025.
11. Huang T, Zhang Y, Niu Y, Xiao Y, Ge Y, Gao J. The Cytidine N-Acetyltransferase NAT10 Promotes Thalamus Hemorrhage-Induced Central Poststroke Pain by Stabilizing Fn14 Expression in Thalamic Neurons. Mol Neurobiol. 2025;62(3):3276-3292.
12. Zhang H, Luo X, Yang W, et al. YTHDF2 upregulation and subcellular localization dictate CD8 T cell polyfunctionality in anti-tumor immunity. Nat Commun. 2024;15(1):9559. Published 2024 Nov 5.
13. Liu H, Xu L, Yue S, et al. Targeting N4-acetylcytidine suppresses hepatocellular carcinoma progression by repressing eEF2-mediated HMGB2 mRNA translation. Cancer Commun (Lond). 2024;44(9):1018-1041.
14. Yin L, Jiang N, Xiong W, et al. METTL16 is Required for Meiotic Sex Chromosome Inactivation and DSB Formation and Recombination during Male Meiosis. Adv Sci (Weinh). 2025;12(3):e2406332.
15. Xiao H, Zhao R, Meng W, Liao Y. Effects and translatomics characteristics of a small-molecule inhibitor of METTL3 against non-small cell lung cancer. J Pharm Anal. 2023;13(6):625-639. doi:10.1016/j.jpha.2023.04.009
16. Weng H, Huang F, Yu Z,et al.The m6A reader IGF2BP2 regulates glutamine metabolism and represents a therapeutic target in acute myeloid leukemia. Cancer Cell. 2022 Dec 12;40(12):1566-1582.e10.
17. Sun YM, Wang WT, Zeng ZC, et al. circMYBL2, a circRNA from MYBL2, regulates FLT3 translation by recruiting PTBP1to promote FLT3-ITD AML progression. Blood. 2019 Oct 31;134(18):1533-1546.
18. ChenH, Gao S, Liu W,et al. RNA N-Methyladenosine Methyltransferase METTL3 Facilitates Colorectal Cancer by Activating the mA-GLUT1-mTORC1 Axis and Is a Therapeutic Target. Gastroenterology 2021 03;160(4).
19. Wei R, Cui X, Min J, et al. NAT10 promotes cell proliferation by acetylating CEP170 mRNA to enhance translation efficiency in multiple myeloma. Acta Pharm Sin B. 2022 Aug; 12(8): 3313–3325.
References
[1] Ingolia NT, Ghaemmaghami S, Newman JR, et al. Genome-wide analysis in vivo of translation with nucleo- tideresolution using ribosome profiling. Science, 2009, 324(5924): 218-223
[2] Gobet C, Naef F. Ribosome profiling and dynamic regulation of translation in mammals. Curr Opin Genet Dev. 2017Apr;43:120-127.
