scRNA-seq
scRNA-seq
Single-cell RNA sequencing
Single-cell RNA sequencing (scRNA-seq) quantifies transcriptomes at single-cell resolution from ultra-low input. EPIBIOTEK® offers scRNA-seq on the MGI DNBelab C-TaiM4 droplet platform, enabling high capture efficiency at reduced library and sequencing cost. The assay supports cell-type–resolved profiling for cancer, stem cell, and developmental studies, including biomarker discovery and translational research.
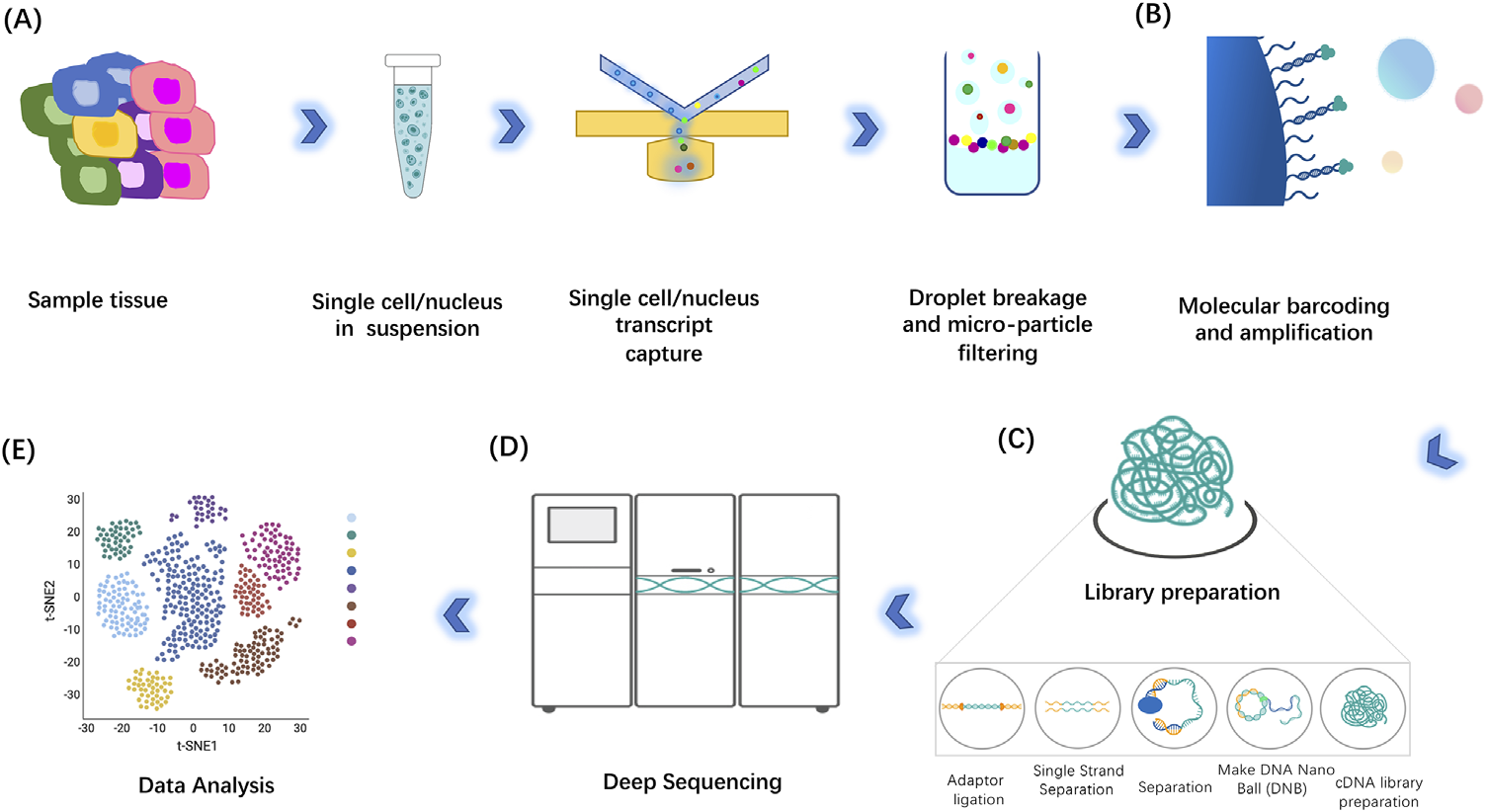
Jovic D, et al. Clin Transl Med. 2022.
Workflow
DNBelab C-TaiM4 uses a Drop-seq–like droplet microfluidics workflow to co-encapsulate individual cells with barcoded gel beads in water-in-oil droplets. After in-droplet lysis, poly(A)+ RNAs are captured by oligo(dT) on cell-barcoded beads and converted to barcoded cDNA by reverse transcription. Following emulsion breakage, sequencing libraries are constructed and sequenced for downstream single-cell analysis. [1]
Advantages
1. Four independent microfluidic channels for flexible processing of 1-4 samples per run
2. High cell-capture efficiency across diverse sample types
3. Lower overall cost with a high fraction of usable reads
Sample Requirements
1. Sample type: cells or tissue-derived cell suspensions (contact regional sales or technical support for detailed requirements)
2. Species: human, mouse, and rat only
Bioinformatic Analysis
Basic analysis
1. Read preprocessing and QC
2. Cell barcode and UMI processing
3. Genome alignment
4. Cell QC and filtering
5. Dimensionality reduction and clustering
6. Automated cell-type annotation
7. Cell-cycle scoring
Advanced analysis
1. Marker-gene functional enrichment (GO, KEGG)
2. Trajectory (pseudotime) inference
3. Cell-cell communication analysis
