Single-Cell CUT&Tag sequencing
nano-CT
Single-Cell CUT&Tag sequencing
、With the rapid advancement of single-cell sequencing technologies in recent years, numerous epigenomic sequencing approaches have progressively evolved to the single-cell level, injecting powerful new momentum into epigenetic research. EPIBIOTEK® officially launches nano-CT, a single-cell CUT&Tag sequencing technology built on an innovative nanobody–Tn5 fusion protein (nano-Tn5) and microfluidic platform. nano-CT offers high sensitivity, flexible modality selection, and high resolution, enabling precise analysis of chromatin states and transcription factor binding at single-cell resolution. Beyond revealing differences across cell types, nano-CT can dissect intra-cell-type heterogeneity, achieving a higher-precision epigenetic landscape. As a powerful new tool, nano-CT is applicable to developmental biology, neuroscience, tumor heterogeneity analysis, clinical translational research, and epigenetic regulatory mechanism studies.
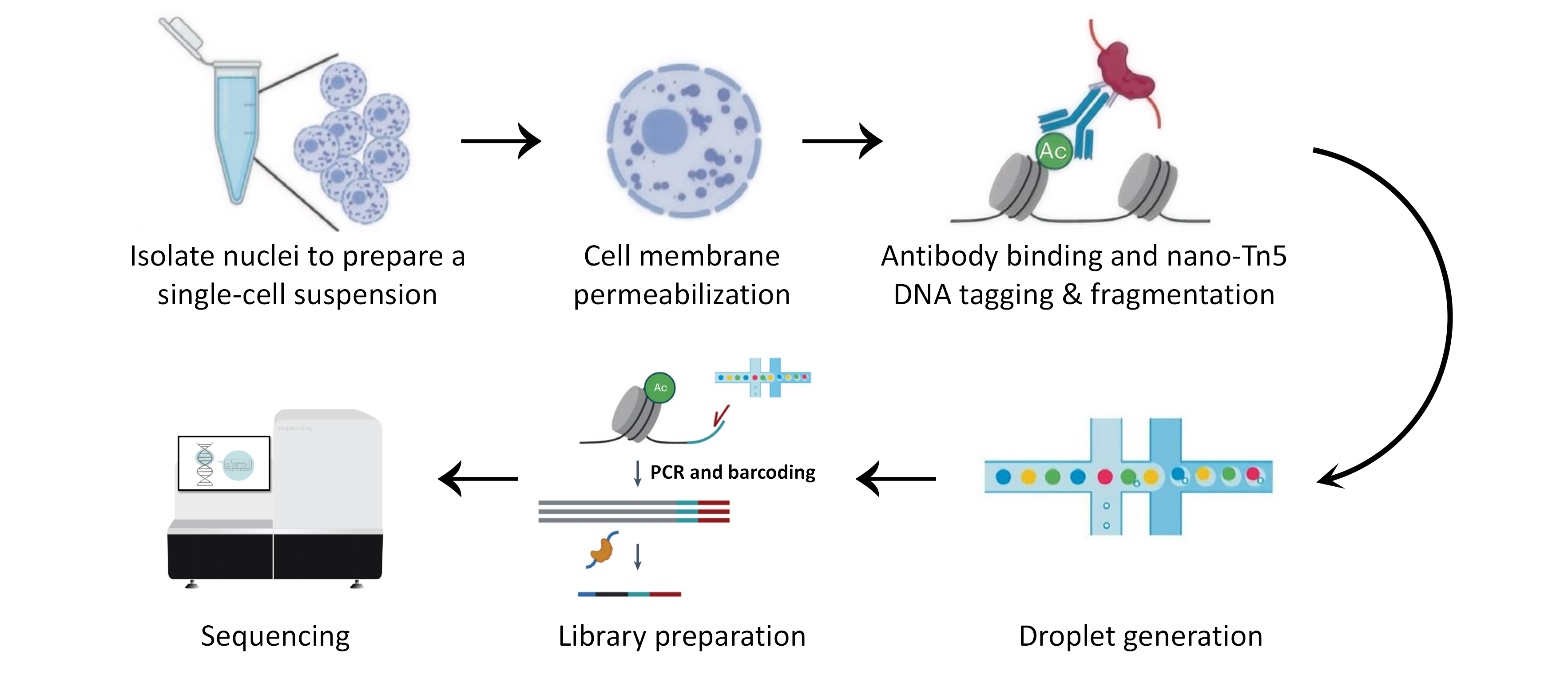
Nuclei extraction and single-cell suspension preparation → Cell membrane permeabilization → Addition of target protein antibody and nanobody-Tn5 fusion protein → DNA tagmentation and fragmentation → Droplet generation → PCR amplification of DNA fragments → nano-CT library construction → High-throughput sequencing → Bioinformatics analysis
Workflow
Advantages
1.Higher sensitivity and fragment recovery with low background noise, improving detection of rare cell populations
2.Supports multimodal detection for deeper understanding of gene regulatory networks
3.Innovative nanobody-Tn5 design eliminates the need for secondary antibodies, simplifying the workflow and improving capture efficiency
Applications
1.Cell Fate and Differentiation Regulation:
Reveal chromatin state differences across cell subpopulations to investigate cell fate determination, functional differentiation, and embryonic development.
2.Tumor Heterogeneity and Therapeutic Strategies:
Analyze chromatin heterogeneity in tumor cells, identify tumor biomarkers, study oncogenic mechanisms, and support the development of new therapeutic strategies.
3.Immune and Nervous System Regulation:
Characterize chromatin landscapes in immune cells, neurons, and glial cells to investigate immune responses, immune-related diseases, and regulatory mechanisms underlying nervous system development and function.
4.Epigenetic Modifications and Gene Regulation:
Study the role of histone modifications in gene regulation, identify transcription factor binding sites, and characterize gene regulatory networks.
5.Multi-Omics Integration and Network Construction:
Combine multi-omics analyses or integrate single-cell multi-omics data to construct multilayered, multidimensional gene regulatory networks.
Sample Requirements
Cells
1.3×10⁵–4×10⁵ cells, viability >85%, cell size 5–40 μm, cell aggregation rate <10%, with no visible debris or contaminants in suspension
Tissue
≥0.1 g/sample
Bioinformatic Analysis
Basic Analysis
1.Data overview
2.Data processing and quality control
3.Peak annotation
4.Valid cell count
5.Peak-Cell Barcode matrix
6.Insert fragment statistics
7.Library complexity analysis
8.Cell clustering and visualization
9.Functional enrichment analysis of marker genes
Advanced Analysis
1.GO and KEGG enrichment analysis of histone modification target genes across groups
2.Motif analysis
3.Trajectory analysis
Custom Analysis
1.Differential histone modification analysis across groups
2.Other (customized)
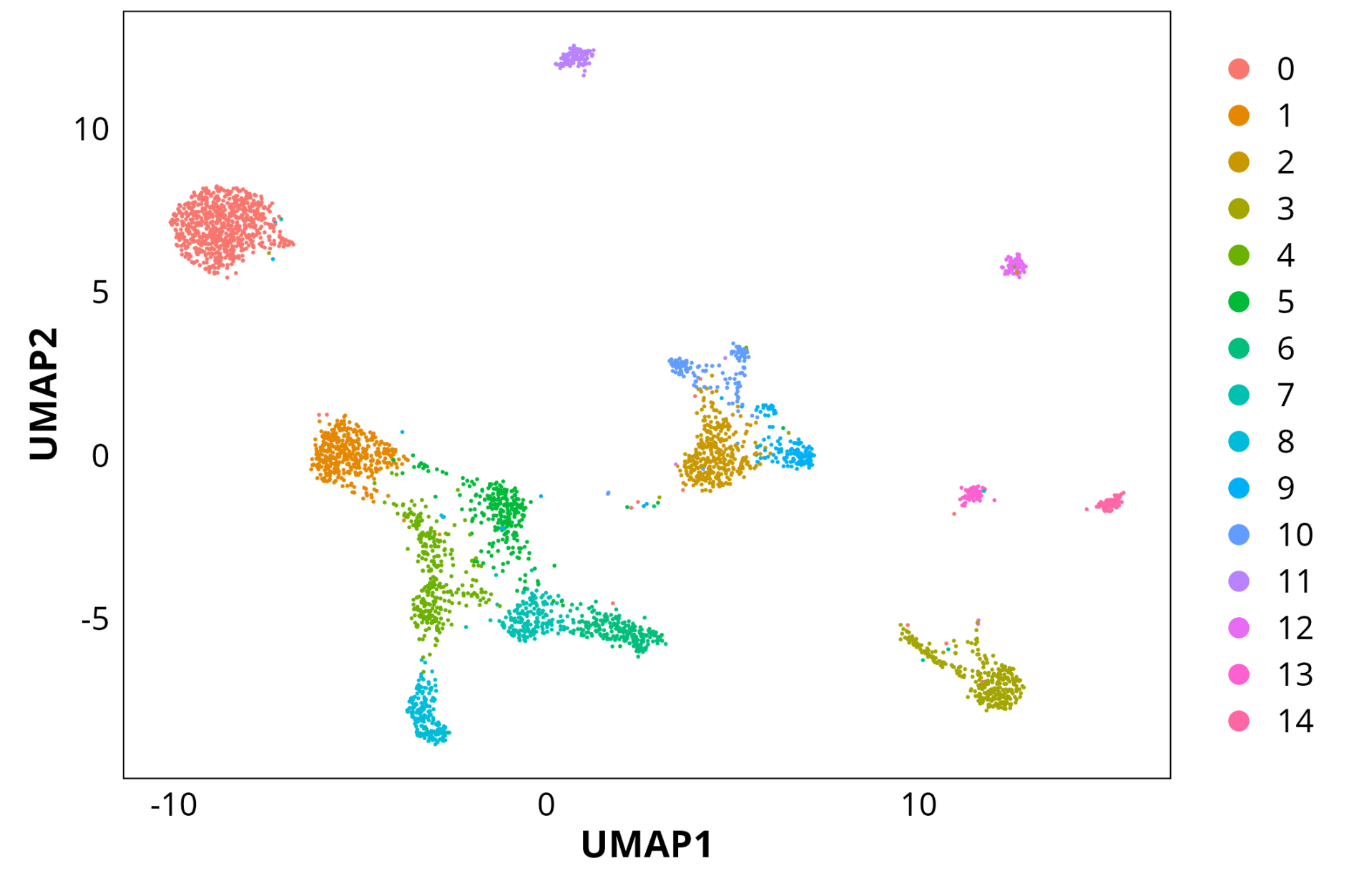
UMAP of cell clusters
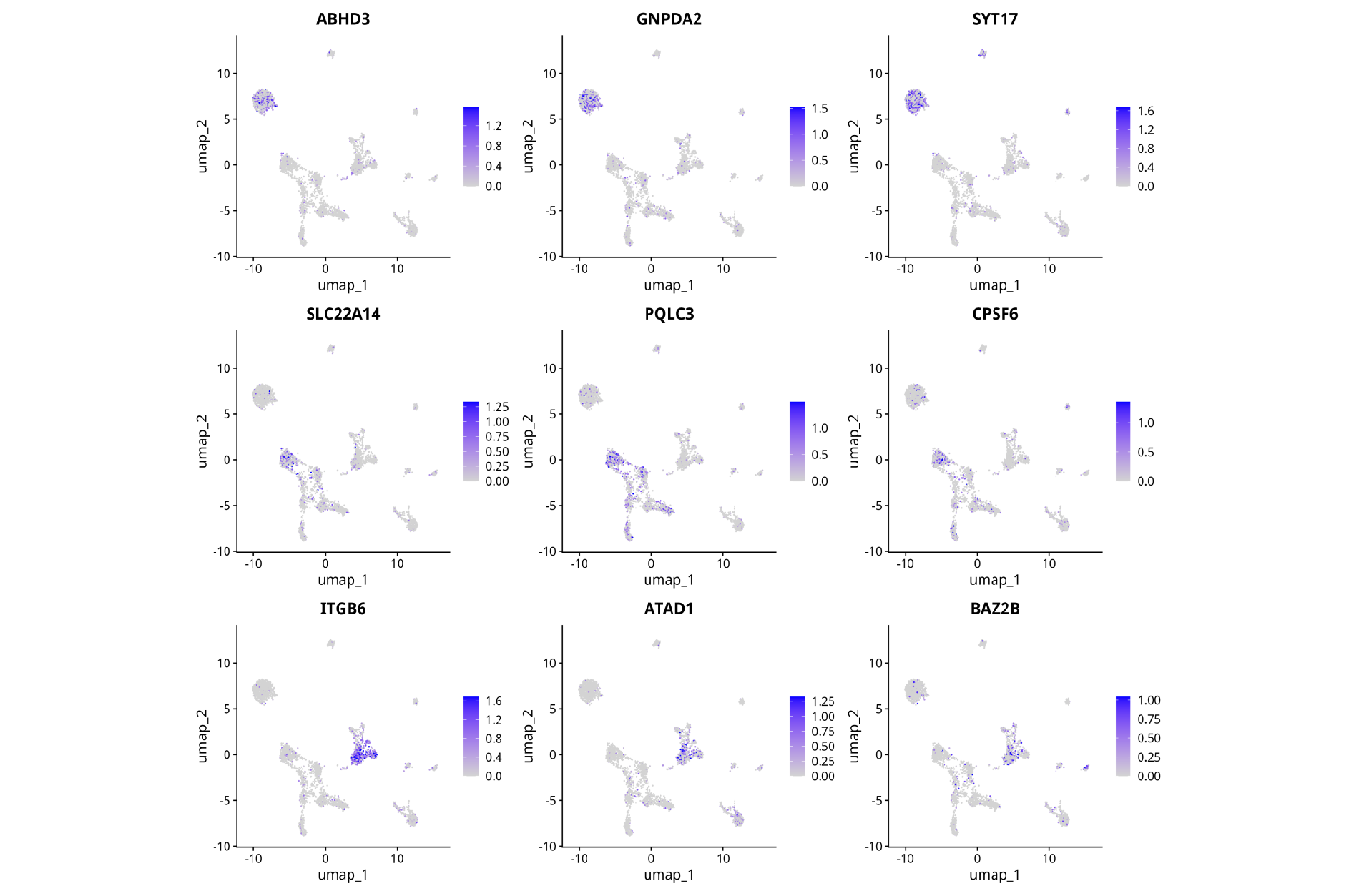
UMAP of cell clusters
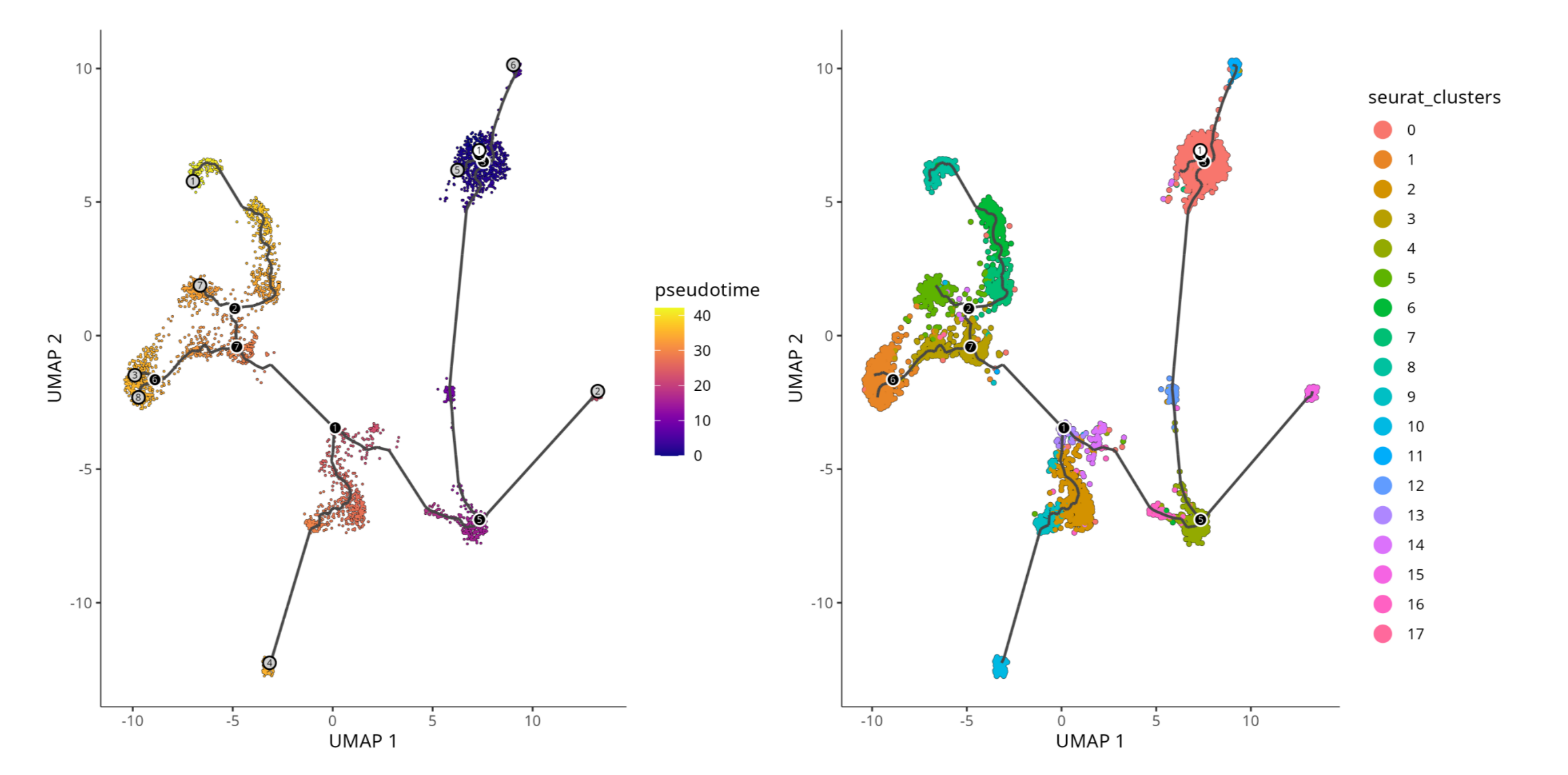
UMAP of trajectory analysis
