acRIP-seq
acRIP-seq
RNA ac4C Modification Sequencing
N4-acetylcytidine (ac4C), an N4-acetylated cytidine, is a conserved chemical modification found in both eukaryotes and prokaryotes. Early studies suggested that ac4C was primarily present on tRNA and 18S rRNA. However, recent research has revealed that ac4C is also abundant on mRNA, where it plays a crucial role in promoting protein translation, influencing RNA stability and alternative splicing, and regulating gene expression. This modification is poised to become an emerging direction in epitranscriptomics.
ac4C immunoprecipitation sequencing (acRIP-seq) comprehensively analyze the distribution and changes of ac4C in the transcriptome, and help researchers to deeply understand the role of RNA acetylation in life and disease, so as to promote the research of precision diagnosis and treatment of diseases.
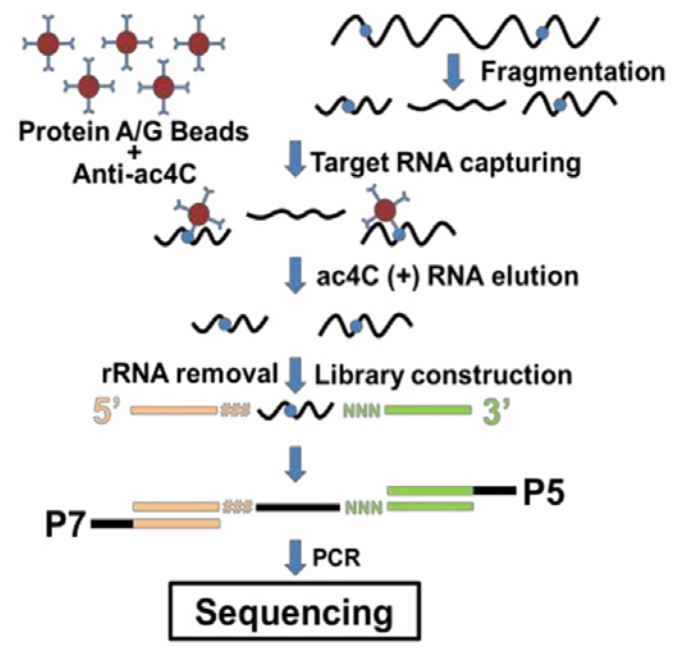
Workflow
acRIP leverages the high specificity of antibodies for acetylated bases to enrich RNA fragments bearing this modification via immunoprecipitation. Subsequent high-throughput sequencing enables genome-wide profiling of acetylated RNA regions across the entire transcriptome, yielding comprehensive and efficient results.
Sample Requirements
>300 μg
Total RNA
>6×10^7
Cells
>150 mg
Tissue
Data Analysis
Basic Analysis:
1. Adapter trimming and QC of raw reads
2. Reference genome alignment
3. Peak Calling
4. Peak annotation
5. Modification analysis of lncRNA and mRNA
6. Metagene plots and pie charts of enriched regions
7. Motif analysis (CXX repeat motif)
8. GO and KEGG analysis of peak-associated genes
9. Differential peak analysis (differential acetylation analysis)
10. GO and KEGG analysis of genes with differential peaks
11. Single-site modification prediction analysis (Single Site, prediction of CXXCXXCXXCXX)
12. Correlation analysis (integration with RNA-seq data)
13. Four-quadrant plot of correlation analysis
14. Heatmap of correlation analysis (only for samples with biological replicates)
Advanced Analysis:
1. Correlation analysis between RNA-seq expression differences and ac4C modification differences
2. Correlation analysis between mRNA alternative splicing and ac4C modification differences
3. Correlation analysis between Ribo-seq translation differences and ac4C modification differences
Demo
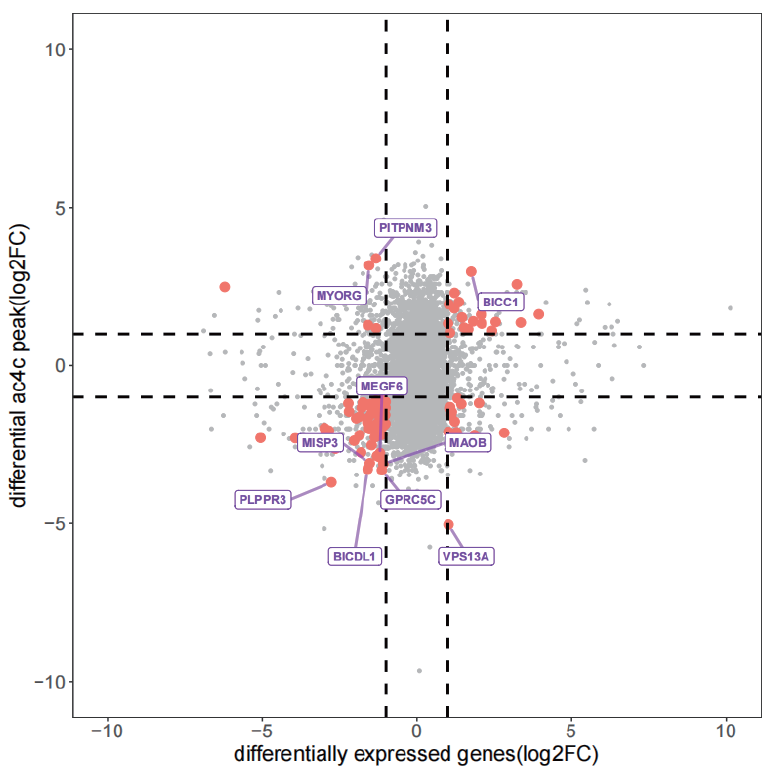
Multimodal correlation analysis four-quadrant plot (RNA-seq and acRIP-seq)
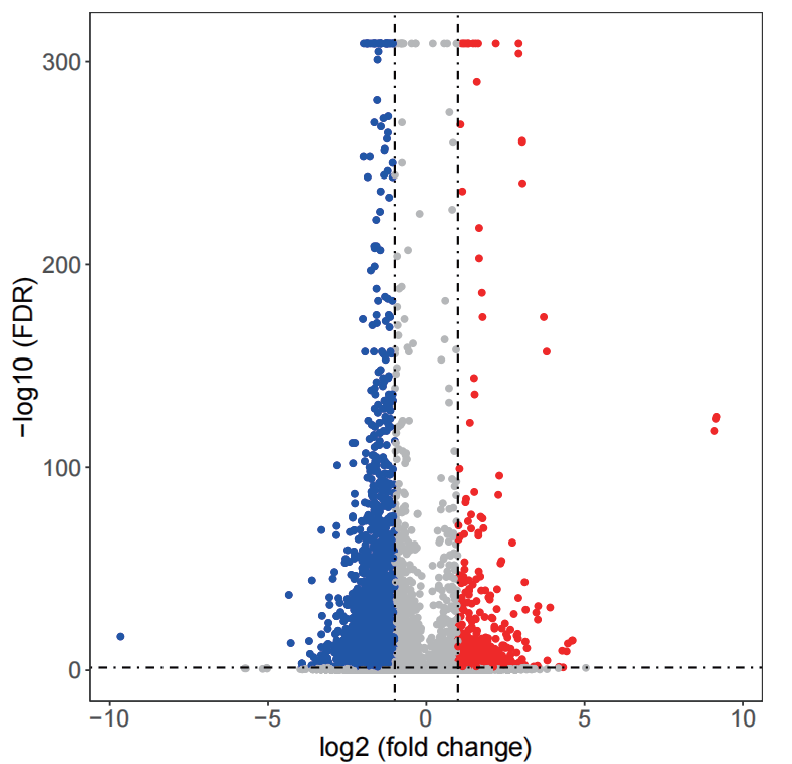
Differences Peaks volcano chart
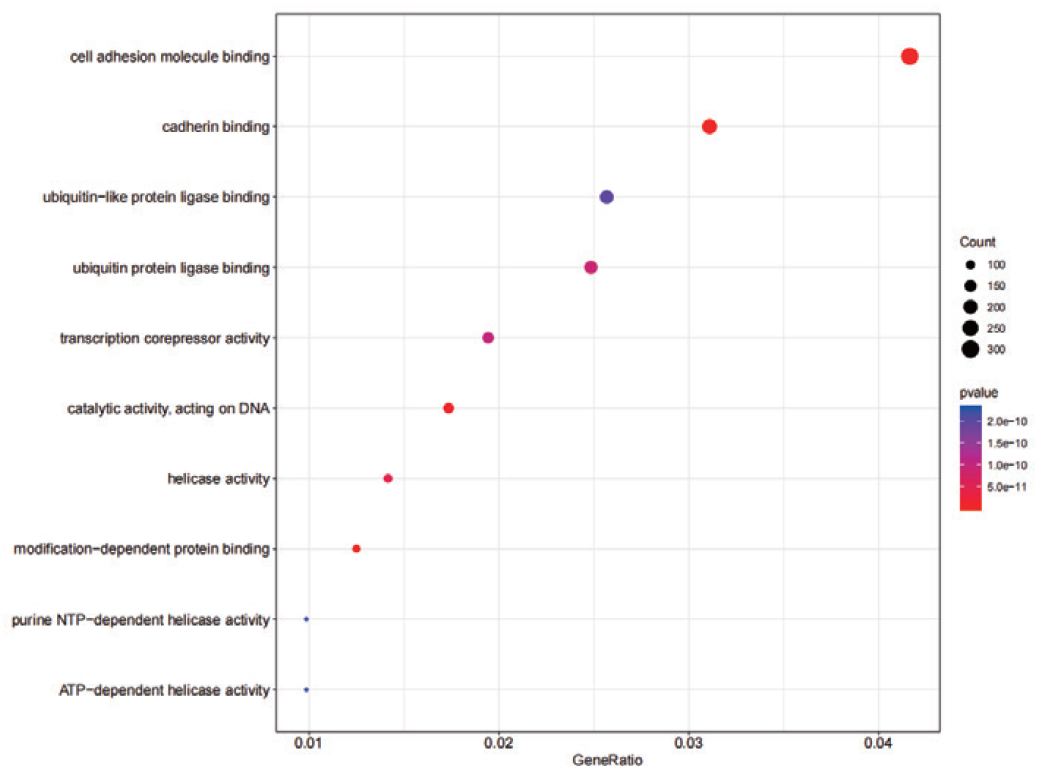
Differential GO analysis bubble plot
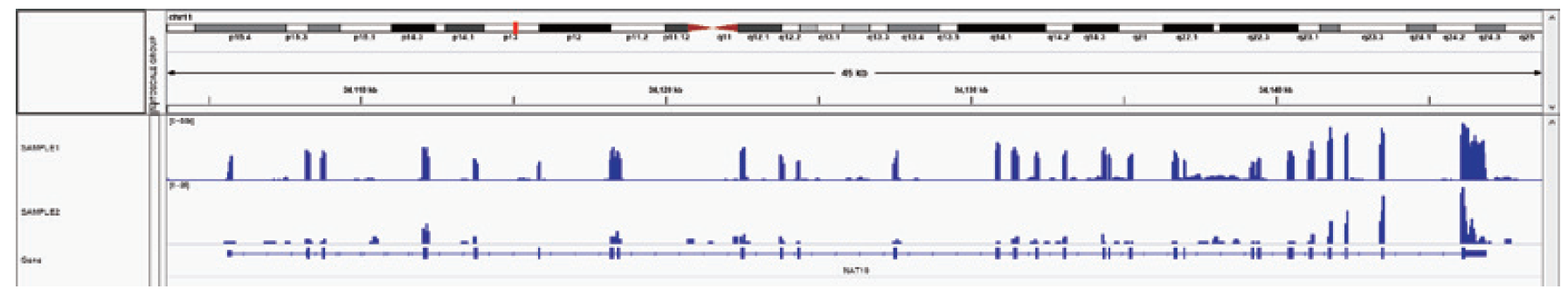
IGV peak plots
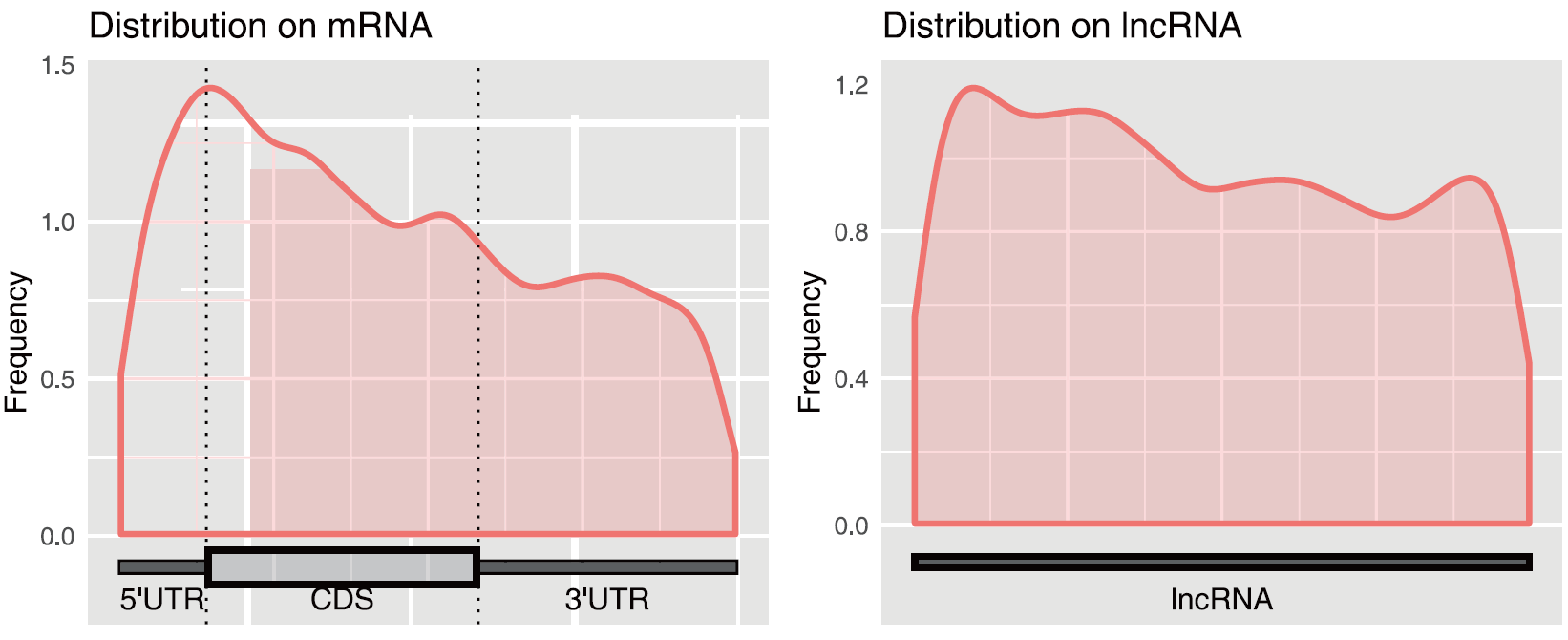
ac4C modified metagene diagram
EPIBIOTEK® Project Publications
1. Zhan YJ, Deng CM, Tang L, et al. NAT10 Increases Lysosomal Acidification to Promote Esophageal Cancer Metastasis via ac4C Acetylation of ATP6V0E1 mRNA. Adv Sci (Weinh). 2025;12(31):e02931.
2. Lei X, Zheng B, Peng Y, et al. Targeting the NAT10/XIST/YAP1 Axis-Mediated Vascular Abnormalization Enhances Immune Checkpoint Blockade in Gastric Cancer. Int J Biol Sci. 2025;21(11):4997-5014. Published 2025 Jul 28.
3. Liu WC, Wei YH, Chen JF, et al. Inhibition of tumor-intrinsic NAT10 enhances antitumor immunity by triggering type I interferon response via MYC/CDK2/DNMT1 pathway. Nat Commun. 2025;16(1):5154. Published 2025 Jun 3.
4. Dai Y, Li Y, Yang X, et al. Silencing the fibronectin gene (FN1) improves NaCl-induced cardiac fibrosis via ferritinophagy-mediated ferroptosis in a nuclear receptor coactivator 4 (NCOA4)-dependent manner. Br J Pharmacol. 2026;183(3):581-597.
5. Wan X, Wang L, Khan MA, et al. NAT10-mediated N4-acetylcytidine modification in KLF9 mRNA promotes adipogenesis. Cell Death Differ. 2025;32(9):1613-1629.
6. Wang L, Chen E, Zhang S, et al. Targeting the ac4C 'Writer' NAT10 enhances pancreatic cancer immunotherapy via dual modulation of CD8+ T cells and tumor cells. Cell Death Dis. 2025;16(1):809. Published 2025 Nov 7.
7. Xiao Z, Wei X, Li P, et al. Impact of N-acetyltransferase 10 on macrophage activation and inflammation-induced cardiac dysfunction. Cell Death Dis. 2025;16(1):471. Published 2025 Jul 1.
8. Li H, Wu H, Li S, et al. Mechanistic study of N-acetyltransferase 10 deficiency enhancing olaparib sensitivity in triple negative breast cancer by inhibiting RAD51 N4-acetylcytidine modification. iScience. 2025;28(7):112860. Published 2025 Jun 9.
9. Song A, Liu B, Li W, et al. Competitive binding between DDX21 and SIRT7 enhances NAT10-mediated ac4C modification to promote colorectal cancer metastasis and angiogenesis- DDX21 promotes colorectal cancer metastasis. Cell Death Dis. 2025;16(1):353. Published 2025 Apr 29.
10. Yu C, Chen Y, Luo H, et al. NAT10 promotes vascular remodelling via mRNA ac4C acetylation. Eur Heart J. 2025;46(3):288-304.
11. Huang T, Zhang Y, Niu Y, Xiao Y, Ge Y, Gao J. The Cytidine N-Acetyltransferase NAT10 Promotes Thalamus Hemorrhage-Induced Central Poststroke Pain by Stabilizing Fn14 Expression in Thalamic Neurons. Mol Neurobiol. 2025;62(3):3276-3292.
12. Xie H, Zhang K, Yin H, et al. Acetyltransferase NAT10 inhibits T-cell immunity and promotes nasopharyngeal carcinoma progression through DDX5/HMGB1 axis. J Immunother Cancer. 2025;13(2):e010301. Published 2025 Feb 12.
13. Zhang X, Zheng Y, Yang J, et al. Abnormal ac4C modification in metabolic dysfunction associated steatotic liver cells. Sci Rep. 2025;15(1):1013. Published 2025 Jan 6.
14. Li KJ, Hong Y, Yu YZ, et al. NAT10 Promotes Prostate Cancer Growth and Metastasis by Acetylating mRNAs of HMGA1 and KRT8. Adv Sci (Weinh). 2024;11(32):e2310131.
15. Xie R, Cheng L, Huang M, et al. NAT10 drives cisplatin chemoresistance by enhancing ac4C-associated DNA repair in bladder cancer. Cancer Res 2023 Mar 20.[2] Yang Q, Lei X, He J, et al. N4-Acetylcytidine Drives Glycolysis Addiction in Gastric Cancer via NAT10/SEPT9/HIF-1α Positive Feedback Loop. Adv Sci (Weinh). 2023 Aug;10(23):e2300898.
16. Shi J, Yang C, Zhang J, et al. NAT10 Is Involved in Cardiac Remodeling Through ac4C-Mediated Transcriptomic Regulation. Circ Res. 2023 Dec 8;133(12):989-1002.
17. Yan Q, Zhou J, Wang Z, et al. NAT10-dependent N4-acetylcytidine modification mediates PAN RNA stability, KSHV reactivation, and IFI16-related inflammasome activation. Nat Commun. 2023 Oct 10;14(1):6327.
