m5C BS-seq
m5C BS-seq
5-Methylcytosine (m5C) is an RNA methylation modification with relatively low abundance. Nevertheless, research on its role in cancer has recently garnered substantial attention. With the emergence of high-throughput sequencing technologies, various nucleotide modifications, including m5C, can now be precisely profiled throughout the transcriptome. m5C exerts influence on RNA stability, alternative splicing, translation efficiency, and post-transcriptional regulation, thereby affecting gene expression and cellular function. Although there is extensive data regarding m5C in tRNA and rRNA, its function in mRNA remains poorly characterized, primarily due to the low abundance of m5C sites in mRNA and the absence of efficient purification strategies (1).
To advance research on mRNA m5C modification, EPIBIOTEK® has recently introduced a single-base resolution mRNA m5C bisulfite sequencing technology service. m5C RNA bisulfite sequencing (m5C-BS-seq) is regarded as the gold standard for m5C mapping, as bisulfite reagents are cost-effective and non-hazardous. Moreover, PCR amplification and sequencing following bisulfite treatment ensure high sensitivity without the need for complex procedures.
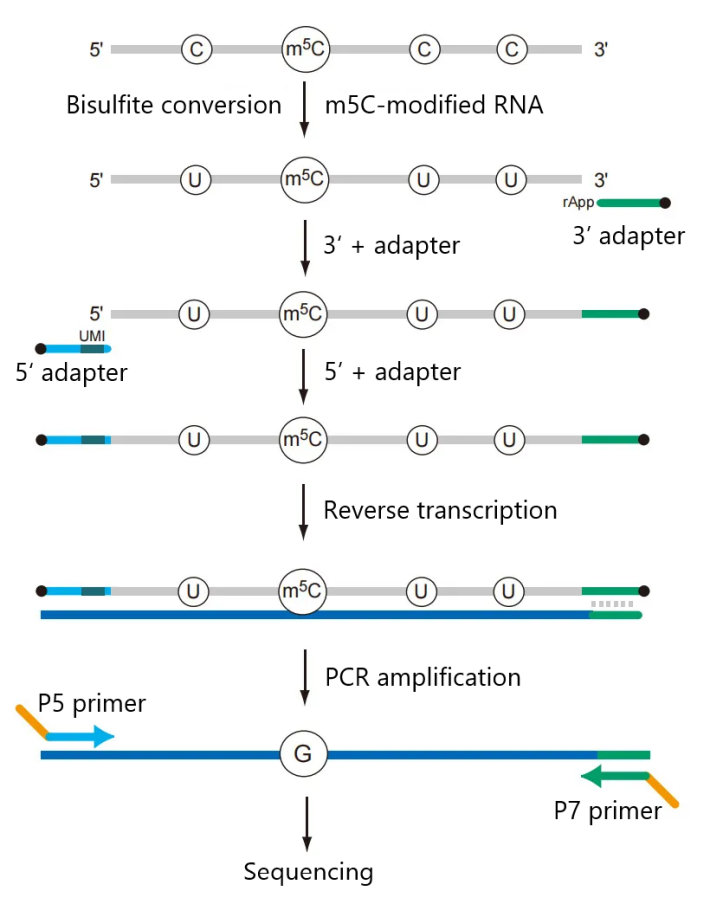
Workflow
Bioinformatics Analysis
m5C BS-seq
1. Raw data quality control and filtering
2. Genome mapping (Alignment)
3. PeakCalling
4. PeakAnno
5. GO enrichment analysis
6. KEGG enrichment analysis
7. Motif analysis
8. Signal matrix analysis (Peak heatmap)
9. CIMS analysis
10. Visualization files
11. Differential analysis
mRNA
1. Raw reads trimming and quality control
2. Genome mapping (Alignment)
3. Gene expression quantification analysis
4. Gene annotation
5. Differential expression analysis
6. Differential gene heatmap plotting (only for biological replicates)
7. GO analysis of differential expressed gene
8. KEGG analysis of differential expressed gene
Sample Requirements
Limited to human, mouse, and rat cells, tissues, and total RNA; other species require evaluation
1. Cells, ≥ 1.5×10⁷ cells/sample
2. Tissues, ≥ 25 mg/sample
3. Total RNA, 50 μg/sample
