GlycoRNA-seq
GlycoRNA-seq
GlycoRNA (glycosylated RNA) is a newly discovered post-transcriptional modification first reported in 2021 by the laboratories of Professors Carolyn Bertozzi and Ryan Flynn at Stanford University. Predominantly localized on the cell surface, GlycoRNA comprises a diverse repertoire of small non-coding RNAs (sncRNAs)—including YRNAs, snRNAs, snoRNAs, small tRNAs, and small rRNAs—that are covalently conjugated with complex N-glycan chainsy (1). Functional studies have demonstrated that GlycoRNA participates in key biological processes, most notably immune regulation; for instance, it can direct neutrophil chemotaxis to counteract infection (2).
EPIBIOTEK® now introduces the first commercial GlycoRNA-seq service, offering researchers a cutting-edge tool to explore this frontier. Our GlycoRNA-seq workflow employs metabolic labeling with Ac4ManNAz followed by click-chemistry enrichment, or the newly developed rPAL method (3), to achieve highly efficient GlycoRNA capture. Coupled with the specialized PANDORA-seq platform (4), the service detects a broader spectrum of sncRNAs bearing unique glycan modifications. Partner with Epibiotek™ to unravel the mysteries of GlycoRNA and open new horizons in RNA modification research.
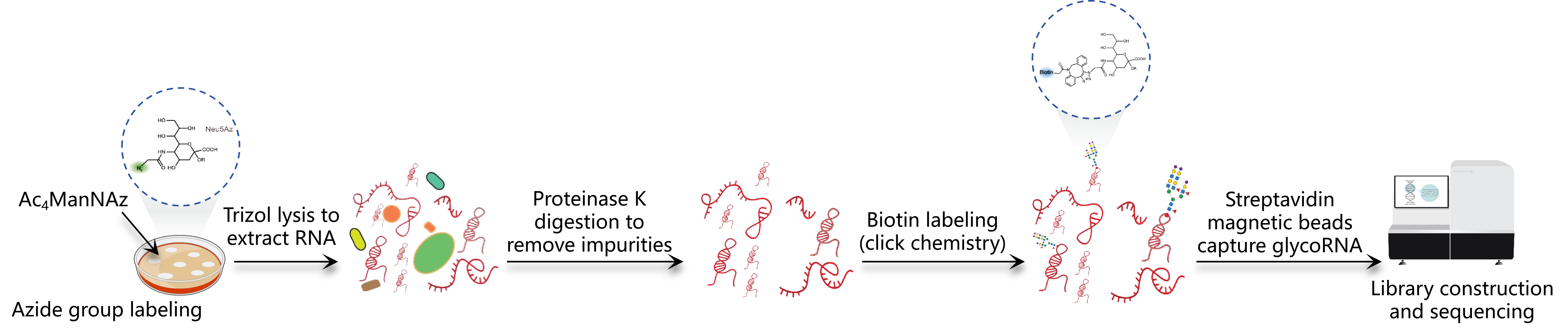
Figure 1. Workflow of Ac4ManNAz method (1,2)
Tips:This method depends on the metabolic activity of the cell and is suitable for living cell samples, not tissue samples
Technical Principles
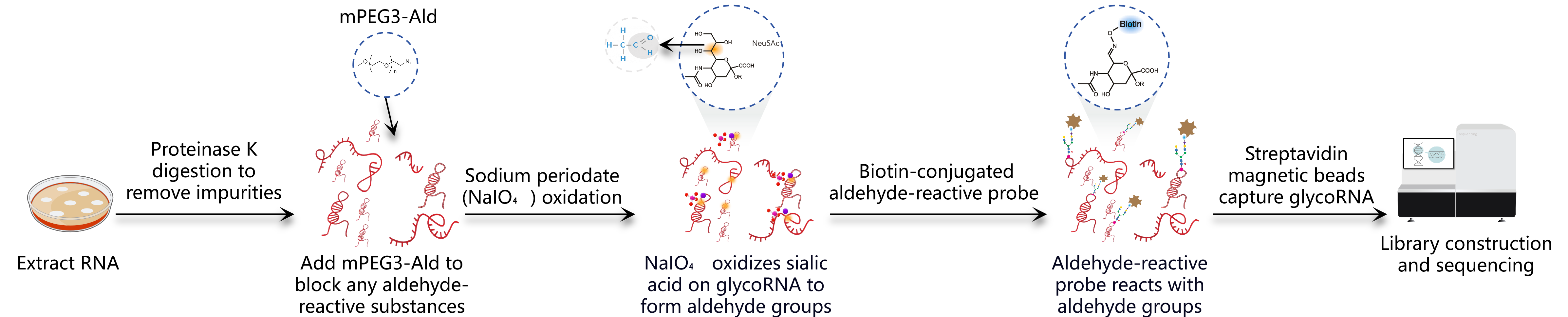
Figure 2. Workflow of rPAL method (3)
Tips: This method is more applicable to a wider range of sample types and has higher capture efficiency
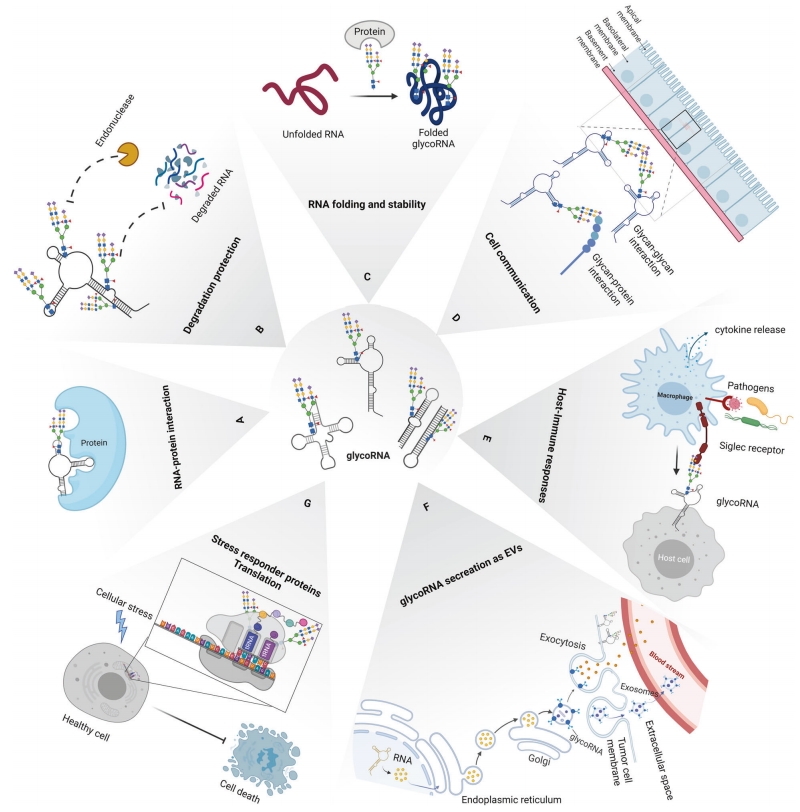
Applications
1. Map GlycoRNA expression profiles, functional variations, and modification patterns across species, tissues, and cell types.
2. Elucidate the regulatory mechanisms of GlycoRNA and dissect its molecular interactions with RNA-binding proteins, lectins, and membrane receptors.
3. Evaluate drug-induced alterations in GlycoRNA abundance and glycosylation to facilitate the development of GlycoRNA-targeted therapeutics.
4. Trace the biogenesis pathway of GlycoRNA and decipher its roles in intercellular communication and cell–cell recognition.
5. Identify disease-associated GlycoRNA biomarkers and uncover their potential as diagnostic or therapeutic targets.
6. Integrate mass-spectrometric analyses to characterize post-translational modifications, determine glycan compositions and linkages, pinpoint glycosylation sites on RNA, and assess co-occurring protein glycosylation—providing an integrated view of GlycoRNA expression, modification, and function.
Advantages
1. Ultra-sensitive detection of low-abundance glycoRNA, delivering comprehensive and high-resolution glycoRNA maps.
2. Simultaneous capture and profiling of diverse sncRNA classes bearing distinct glycan modifications.
3. Rich downstream analyses encompassing sequence annotation, differential expression of each sncRNA subtype, and—when integrated with mass-spectrometric data—prediction of glycan classes and structures on glycoRNA, as well as quantitative comparison of glycan compositions across samples.
Sample Requirements
1. Cells, Ac4ManNAz ≥ 3×10⁶ cells/sample,rPAL≥2×10⁶ cells/sample
2. Tissue samples:tissue for RNA later processing, ≥0.2 cm (applicable only to the rPAL method; approximately 30μg total RNA can be extracted from 25 mg tissue).
3. Whole blood: recommended ≥ 5 mL, minimum 3 mL.
4. RNA: Ac4ManNAz :≥50 μg (marked with Ac4ManNAz) rPAL : ≥30μg
5. The recommended sample size is 3 biological replicates per group (3 vs 3) or more replicates.
Sample Types: Limited to human, mouse, and rat cells; other species require evaluation.
Bioinformatic Analysis
Basic Analysis
1. Raw reads trimming and quality control
2. Alignment to the reference genome
3. Non-coding RNA database comparison (tRNA, rRNA, etc.)
4. SncRNA length and classification statistics
5. Quantitative analysis of various sncRNAs
6. Analysis of expression differences of various sncRNAs (tsRNA, rsRNA, piRNA, etc.)
7. Differential expression sncRNA clustering analysis (limited to biological repeats)
Advanced Analysis
1. Customers can select no more than 50 sncRNA for analysis
2. Prediction of target gene
3. GO analysis of target gene
4. KEGG analysis of target genes
Related Articles
Cell:Glycosylated RNA is the key for neutrophil recruitment (2)
This study elucidates the role of GlycoRNA in orchestrating neutrophil recruitment in mice. GlycoRNA is predominantly displayed on neutrophils and is indispensable for their migration to inflammatory foci. Mechanistically, neutrophil GlycoRNA engages P-selectin on endothelial cells, thereby facilitating neutrophil adhesion and transmigration. The Sidt gene family (Sideroflexin transporters) is shown to be critical for GlycoRNA biosynthesis and function; genetic ablation of these genes markedly attenuates neutrophil recruitment. Collectively, the findings unveil a previously unrecognized biological dimension in which RNA-mediated glycosylation governs cellular behavior, and further experiments corroborate GlycoRNA as a key mediator of neutrophil–endothelial interactions.
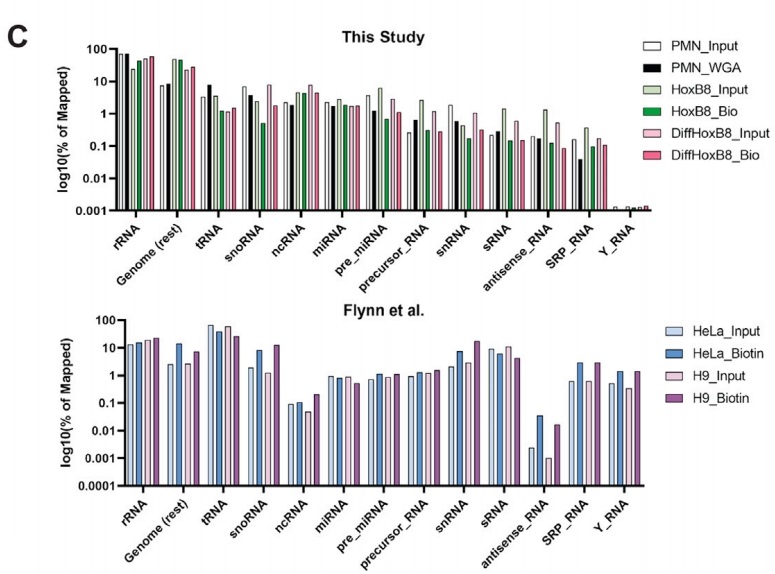
Figure 3. The types of GlycoRNA in different cell types
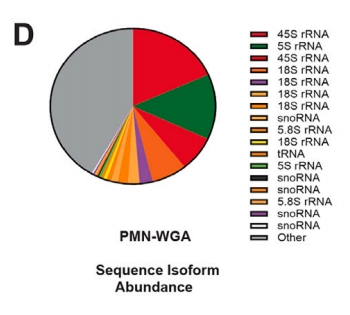
Figure 4. Pie chart of GlycoRNA isoform source ratio
References
[1] Flynn RA, Pedram K, Malaker SA, et al. Small RNAs are modified with N-glycans and displayed on the surface of living cells. Cell. 2021;184(12):3109-3124.e22.
[2] Zhang N, Tang W, Torres L, et al. Cell surface RNAs control neutrophil recruitment. Cell. 2024;187(4):846-860.e17.
[3] Xie Y, Chai P, Till NA, et al. The modified RNA base acp3U is an attachment site for N-glycans in glycoRNA. Cell. 2024;187(19):5228-5237.e12.
[4] Shi J, Zhang Y, Tan D, et al. PANDORA-seq expands the repertoire of regulatory small RNAs by overcoming RNA modifications . Nat Cell Biol. 2021;23(4):424-436.
