MeRIP-seq(Micro Methylated RNA Immunoprecipitation Sequencing)

Introduction
m6A is a prevalent RNA methylation modification. Studies have shown that it plays crucial roles in regulating gene expression, splicing, RNA editing, RNA stability, controlling mRNA lifespan and degradation, and mediating circular RNA translation.
As one of the earliest companies in China to offer MeRIP-seq services, Epibiotek® has been dedicated to optimizing MeRIP technology, enabling library preparation from low-input sample amounts. Our bioinformatics analysis includes popular figures from high-impact publications, supporting researchers in conducting comprehensive and in-depth studies of post-transcriptional m6A modification profiles.
Advantages
1. Low sample requirement, 500 ng~20 μg total RNA is sufficient.
2. The analysis content was upgraded again to include popular charts of high-scoring articles.
3. The experimental technology and bioinformatics analysis technology are excellent, and the motif analysis results guarantee significant RRACH characteristics, such as no full refund or free retest.
Applications
1. Draw a transcriptome-wide m6A modification map.
2. Study the abnormal expression of m6A modification in tumors and other diseases.
3. Investigate the regulatory mechanisms of various physiological/pathological processes that determine cell fate.
Sample Requirements
Limited to human, mouse, and rat cells, tissues, and total RNA; other species require evaluation.
m6A MeRIP-seq
- Total RNA > 10 μg
- Cells > 5×10⁶
- Tissue > 10 mg
PolyA m6A MeRIP-seq
- Total RNA > 5 μg
- Cells > 1.5×10⁷
- Tissue > 50 mg
Bioinformatic Analysis
Basic Analysis
1. Adapter trimming and quality control of raw reads
2. Reference genome mapping
3. Peak calling
4. Peak annotation
5. lncRNA and mRNA modification analysis
6. Metagene plot and pie chart of enriched regions
7. Motif analysis
8. GO and KEGG analysis of peak-associated genes
9. Differential peak analysis (differential modification analysis for lncRNA and mRNA)
10. GO and KEGG analysis of genes with differential peaks
Advanced Analysis
1. Single-site modification prediction analysis
2. Correlation analysis (with RNA-seq or multi-omics integration)
3. Quadrant plot for correlation analysis
4. Heatmap for correlation analysis
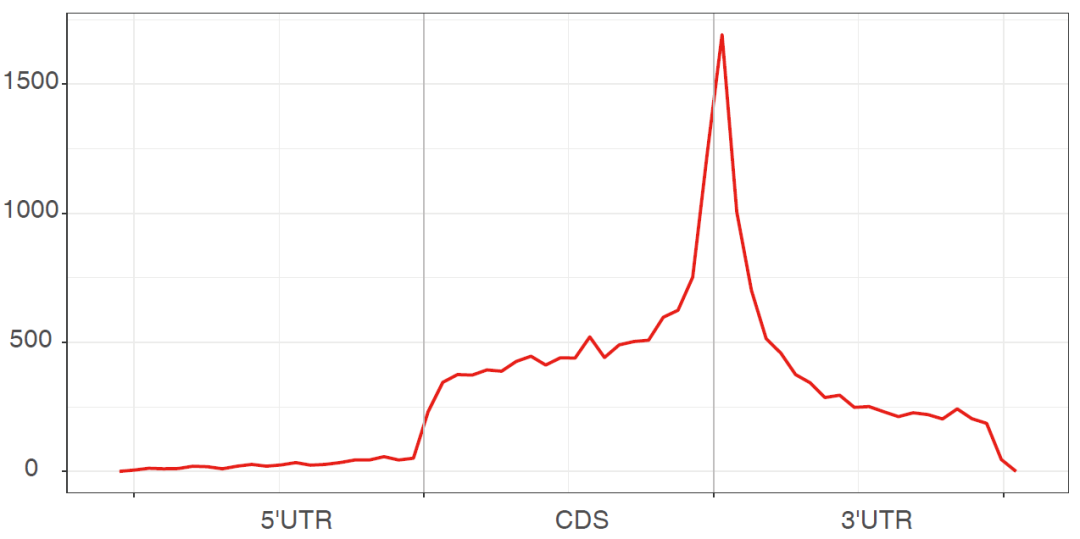
Figure 2. m6A modified metagenes picture
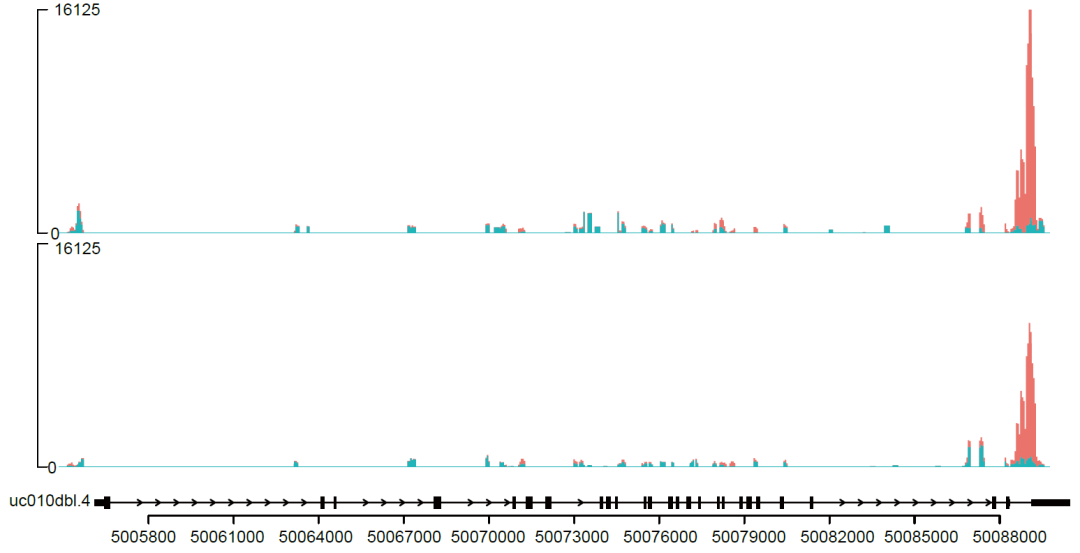
Figure 4. IGV peak plots
Project Publications
Nat Commun:Mettl3-mediated mRNA m6A modification controls postnatal liver development by modulating the transcription factor Hnf4a (1)
Using Alb-Cre-driven Mettl3 conditional knockout (Mettl3-cKO) during the perinatal period, this study demonstrates that loss of Mettl3 triggers extensive hepatocyte apoptosis and steatosis, culminating in severe liver injury and lethality within seven weeks after birth. meRIP-seq (provided by Epibiotek™) combined with RNA-seq revealed that multiple liver-enriched transcription-factor mRNAs—most notably Hnf4a, the master regulator of hepatocyte identity—carry m⁶A modifications. Deletion of Mettl3 reduces m⁶A on Hnf4a mRNA, leading to Igf2bp1-dependent destabilization of the transcript and consequent down-regulation of Hnf4a protein. AAV8-mediated restoration of Hnf4a expression markedly ameliorates liver damage and extends the lifespan of Mettl3-cKO mice. In contrast, tamoxifen-inducible Alb-CreERT2-mediated deletion of Mettl3 in adult mice does not perturb hepatic homeostasis. Collectively, these findings delineate the dynamic and stage-specific requirement for Mettl3-mediated m⁶A RNA modification in mammalian liver development.
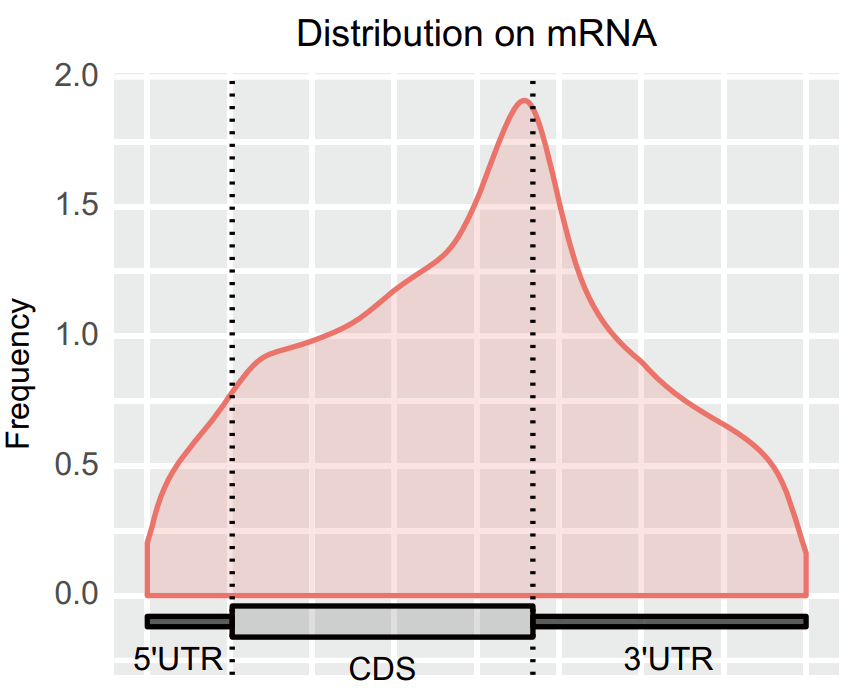
Figure 5. m6A modification of mRNA in mouse liver at week of age metagene picture
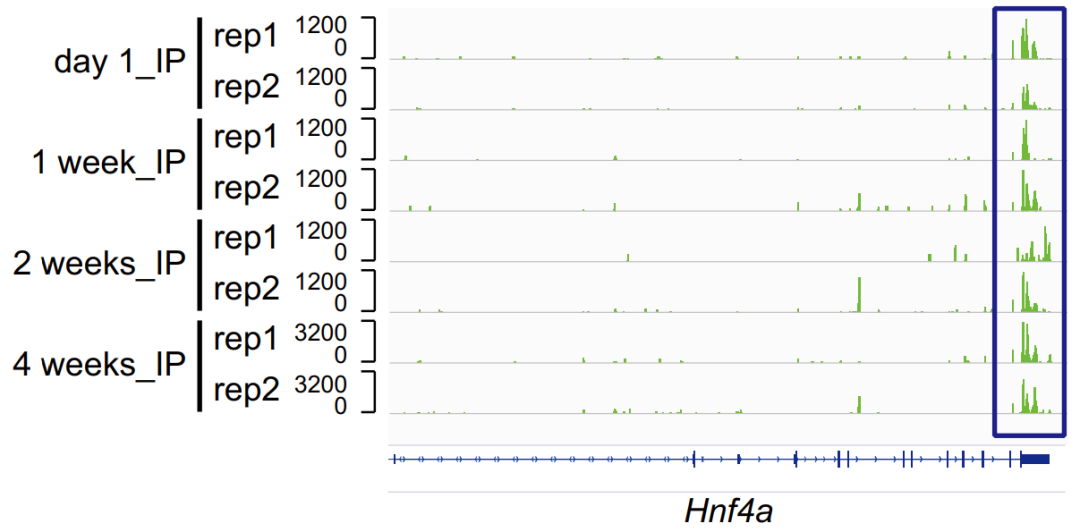
Figure 6. m6A modification peaks at the Hnf4a site in samples taken at four different time points after birth
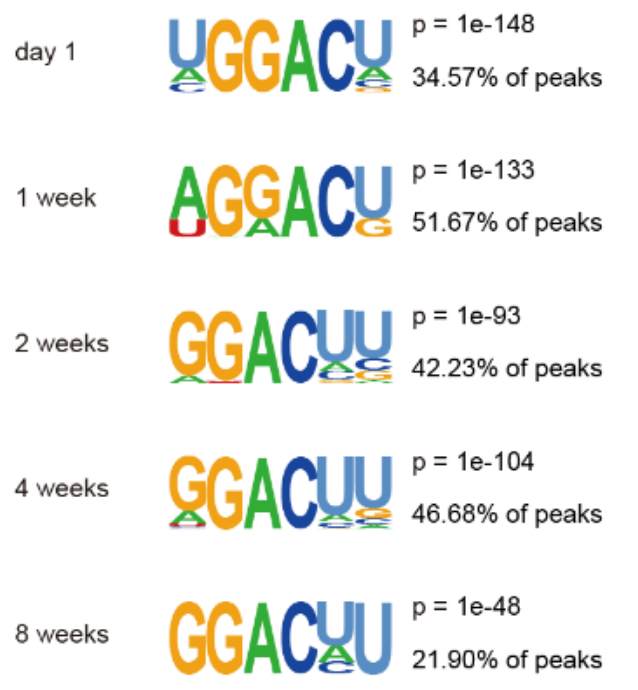

Figure 7. Analysis of the modified Motif m6A
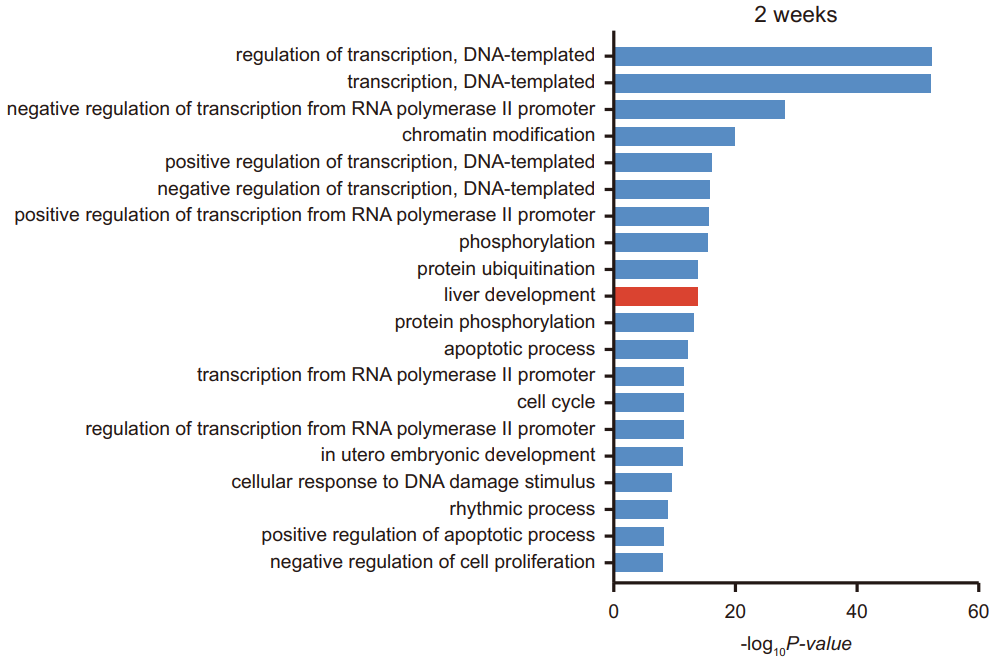
Figure 8. GO enrichment analysis of m6A modified genes in mouse liver tissue at week of age
EPIBIOTEK® Project Publications
[1] Xu Y, Zhou Z, Kang X, et al. Mettl3-mediated mRNA m6A modification controls postnatal liver development by modulating the transcription factor Hnf4a. Nat Commun. 2022 Aug 5;13(1):4555.
[2] Jie Lai, Zhiyong Zhou, Kan Hu, et al. (2024) N6-methyladenosine methylation analysis of long noncodingRNAs and mRNAs in 5-FU-resistant colon cancer cells, Epigenetics, 19:1, 2298058 .
[3] Zhang Y, Yan C, Xie Q, et al. Exposure to bisphenol A affects transcriptome-wide N6-methyladenine methylation in ovarian granulosa cells. Ecotoxicol Environ Saf. 2024 Mar 1;272:116071.
[4] Shi Y, Niu Y, Yuan Y, et al. PRMT3-mediated arginine methylation of IGF2BP1 promotes oxaliplatin resistance in liver cancer. Nat Commun. 2023 Apr 6;14(1):1932.
[5] Xu H, Lin X, Li Z, et al. VIRMA facilitates intrahepatic cholangiocarcinoma progression through epigenetic augmentation of TMED2 and PARD3B mRNA stabilization. J Gastroenterol. 2023 Sep;58(9):925-944.
[6] Li Y, He X, Lu X, et al. METTL3 acetylation impedes cancer metastasis via fine-tuning its nuclear and cytosolic functions. Nat Commun 2022 Oct 26;13(1).
[7] Liu L, Wu Y, Li Q, et al. METTL3 Promotes Tumorigenesis and Metastasis through BMI1 m6A Methylation in Oral Squamous Cell Carcinoma, Molecular Therapy (2020).
