ATAC-seq
ATAC-seq
ATAC-seq maps chromatin accessibility by Tn5-mediated fragmentation and adapter tagging of open chromatin. Standard ATAC-seq requires nuclei isolation prior to tagmentation; this step is laborious and operator dependent, leading to variable membrane removal and reduced consistency in turnaround and success. This product provides a proprietary one-step ATAC buffer that enables direct tagmentation after simple processing of PBS-washed cells, streamlining library preparation. Buffer-treated cells can be stored at 4°C for up to 72 h, improving flexibility for clinical samples.
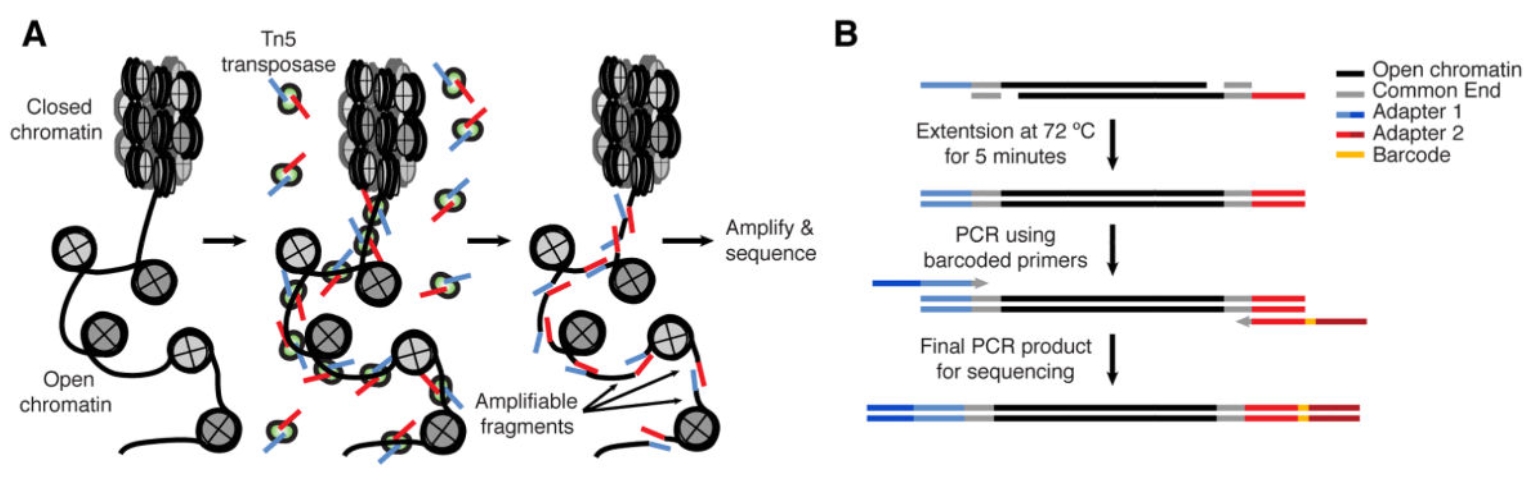
ATAC-seq principle and workflow(Buenrostro JD, et al. Curr Protoc Mol Biol. 2015.)
Workflow
A. Library construction. Tn5 transposase loaded with sequencing adapters preferentially inserts into accessible chromatin; PCR amplification of tagged fragments enables identification of open regions.
B. Post-tagmentation. After 72°C extension to complete adapter incorporation and subsequent amplification, PCR products contain adapters, common sequence ends, and sequencing tags.
Applications
1. Assess whether epigenetic regulation is involved in the biological question of interest
2. Identify regulatory transcription factors via motif analysis
3. Identify TF target genes and functional elements
4. Combine with super-enhancer calling to define active super-enhancer regions
5. Characterize regulatory programs and cellular responses in drug treatment or disease
6. Identify transcription factors associated with cell fate, disease, or response phenotypes
Advantages
1. Low-input, high sensitivity: 50,000 cells/sample
2. Proprietary one-step reagent improves reproducibility and success rate
3. Enables joint inference of open chromatin loci and transcription factor binding-related signals
Sample requirements
Human, mouse, and rat; other species require evaluation
50,000 cells/sample
Live cells
≥ 100 mg
Tissue
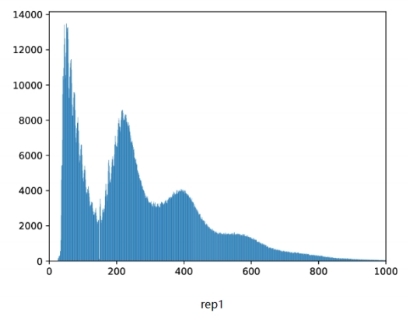
Fragment size distribution
Data Analysis
1.Genome alignment
2.Enriched-region analysis
2.1 Peak calling
2.2 Peak distribution (heatmap, metaProfile)
2.3 GO enrichment of peak-associated genes
2.4 KEGG enrichment of peak-associated genes
2.5 Differential peak analysis
2.6 GO/KEGG enrichment of genes associated with differential peaks
3.Motif analysis of differentially enriched regions
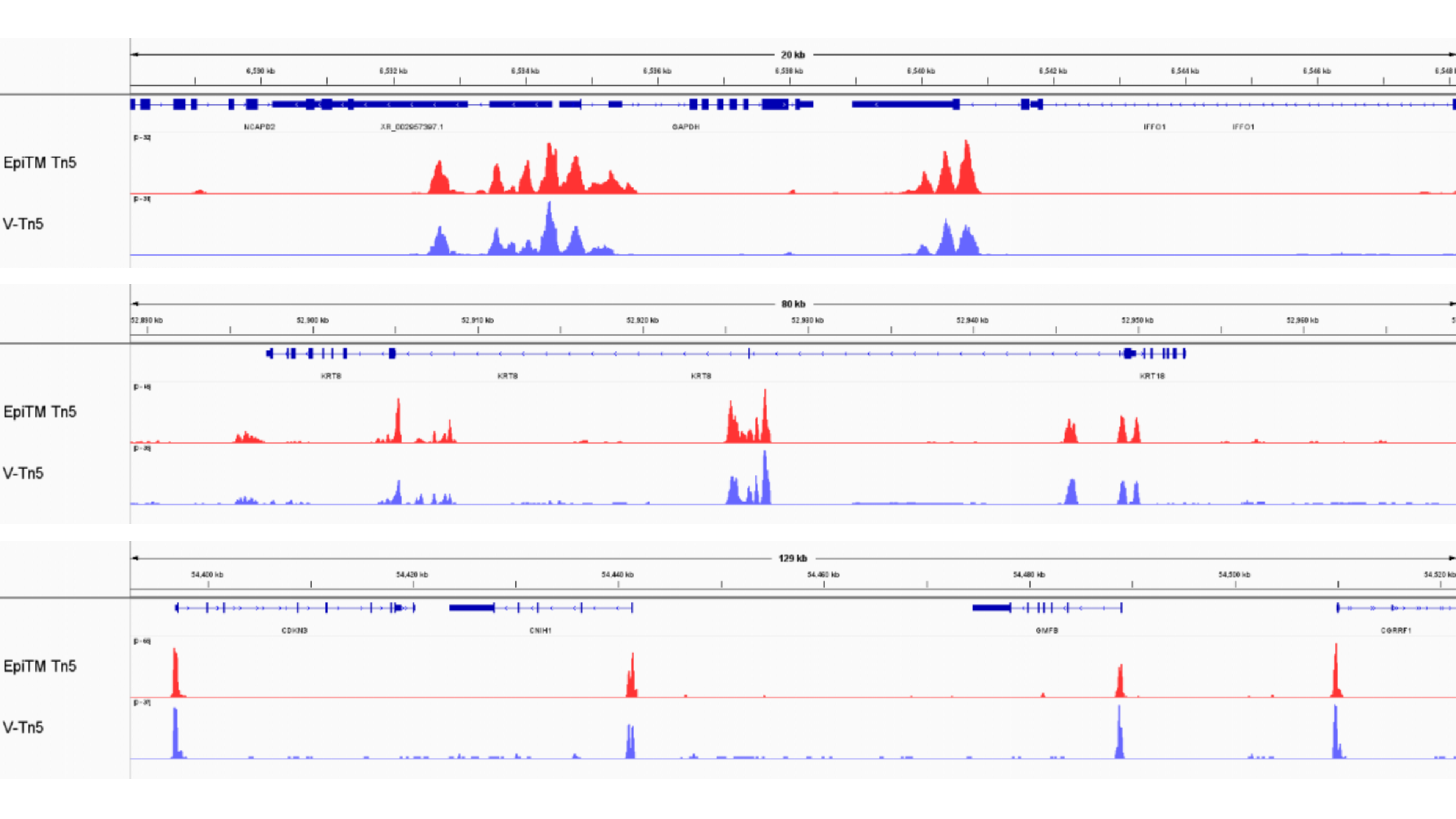
IGV peak tracks (top: EPIBIOTEK data; bottom: V company kit data)
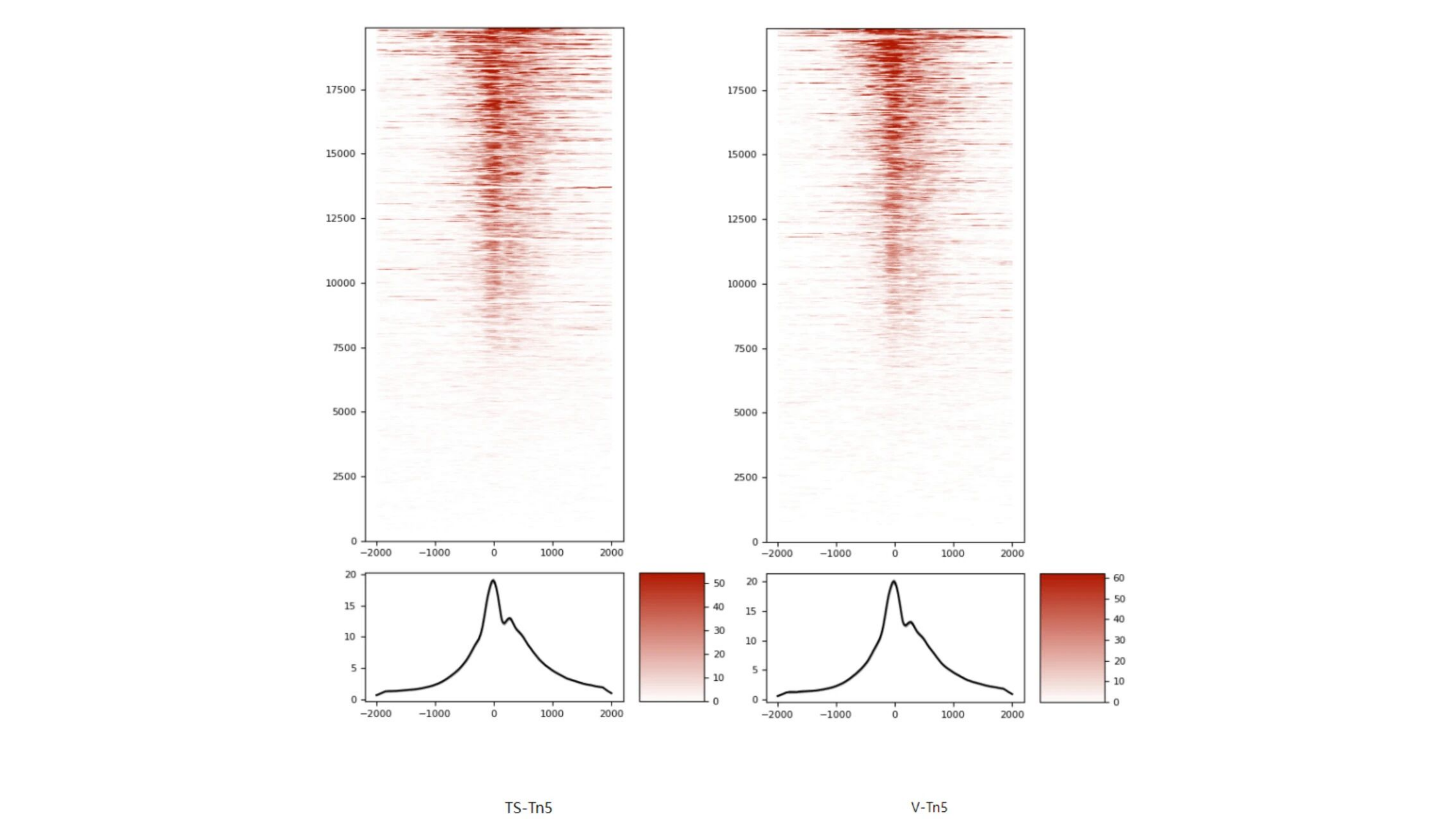
TSS enrichment heatmap (left: EPIBIOTEK data; right: V company kit data)
EPIBIOTEK® Project Publications
1. Wu J,Chen Y, Liao Z, et al. Self-amplifying loop of NF-κB and periostin initiated by PIEZO1 accelerates mechano-induced senescence of nucleus pulposus cells and intervertebral disc degeneration. Mol Ther. 2022 May 26:S1525-0016(22)00319-7.
2. Wang Y, Chen K, Liu G, et al. Disruption of Super-Enhancers in Activated Pancreatic Stellate Cells Facilitates Chemotherapy and Immunotherapy in Pancreatic Cancer. Adv Sci (Weinh). 2024;11(16):e2308637.
3. Chen Y, Zhou T, Liao Z, et al. Hnrnpk is essential for embryonic limb bud development as a transcription activator and a collaborator of insulator protein Ctcf. Cell Death Differ. 2023 Oct;30(10):2293-2308.
4. Xu J, Chen Z, Huang X, et al. Multi-omics analysis reveals that linoleic acid metabolism is associated with variations of trained immunity induced by distinct BCG strains. Sci Adv. 2024 Apr 5;10(14):eadk8093.
5. You B, Xia T, Gu M, et al. AMPK-mTOR-Mediated Activation of Autophagy Promotes Formation of Dormant Polyploid Giant Cancer Cells. Cancer Res 2022 Mar 01;82(5).
6. Qin G, Liu Z, Yang J, et al. Targeting specific DNA G-quadruplexes with CRISPR-guided G-quadruplex-binding proteins and ligands. Nat Cell Biol. 2024;26(7):1212-1224.
7. Zhong C, Li N, Wang S, et al. Targeting osteoblastic 11β-HSD1 to combat high-fat diet-induced bone loss and obesity. Nat Commun. 2024;15(1):8588. Published 2024 Oct 4.
8. Xu JC, Chen ZY, Huang XJ, et al. Multi-omics analysis reveals that linoleic acid metabolism is associated with variations of trained immunity induced by distinct BCG strains. Sci Adv. 2024;10(14):eadk8093.
9. Wang L, Chen E, Zhang S, et al. Targeting the ac4C 'Writer' NAT10 enhances pancreatic cancer immunotherapy via dual modulation of CD8+ T cells and tumor cells. Cell Death Dis. 2025;16(1):809. Published 2025 Nov 7.
10. Su Y, Li S, Wang L, et al. Loss of SETD8 Impairs Mitochondrial Homeostasis to Suppress Leukemia Stem Cell Function in t(8;21) Acute Myeloid Leukemia. Cancer Res. 2025;85(21):4122-4138.
11. Zhang H, Luo X, Yang W, et al. YTHDF2 upregulation and subcellular localization dictate CD8 T cell polyfunctionality in anti-tumor immunity. Nat Commun. 2024;15(1):9559. Published 2024 Nov 5.
12. Wang Y, Chen K, Liu G, et al. Disruption of Super-Enhancers in Activated Pancreatic Stellate Cells Facilitates Chemotherapy and Immunotherapy in Pancreatic Cancer. Adv Sci (Weinh). 2024;11(16):e2308637.
13. Wang WL, Song ZX, Bu SM, et al. Ythdc1-p300-Klf5 Complex-Mediated Golgi Dysfunction Promotes Aortic Aneurysm. Adv Sci (Weinh). 2026;13(4):e12116.
