ACC-seq
ACC-seq
Assay for Chromatin-bound Condensates by exploratory Sequencing (ACC-seq) represents a novel technological innovation developed by the Yinqing Li and Pilong Li research team at Tsinghua University (1), designed for the genome-wide detection of chromatin-bound condensates. EPIBIOTEK® takes pride in presenting this cutting-edge sequencing service, which finds extensive application in research concerning condensate formation mechanisms, transcription factor screening, and the pathogenesis of diseases. ACC-seq allows for the accurate localization of chromatin condensates and promotes the in-depth analysis of gene expression regulatory networks.
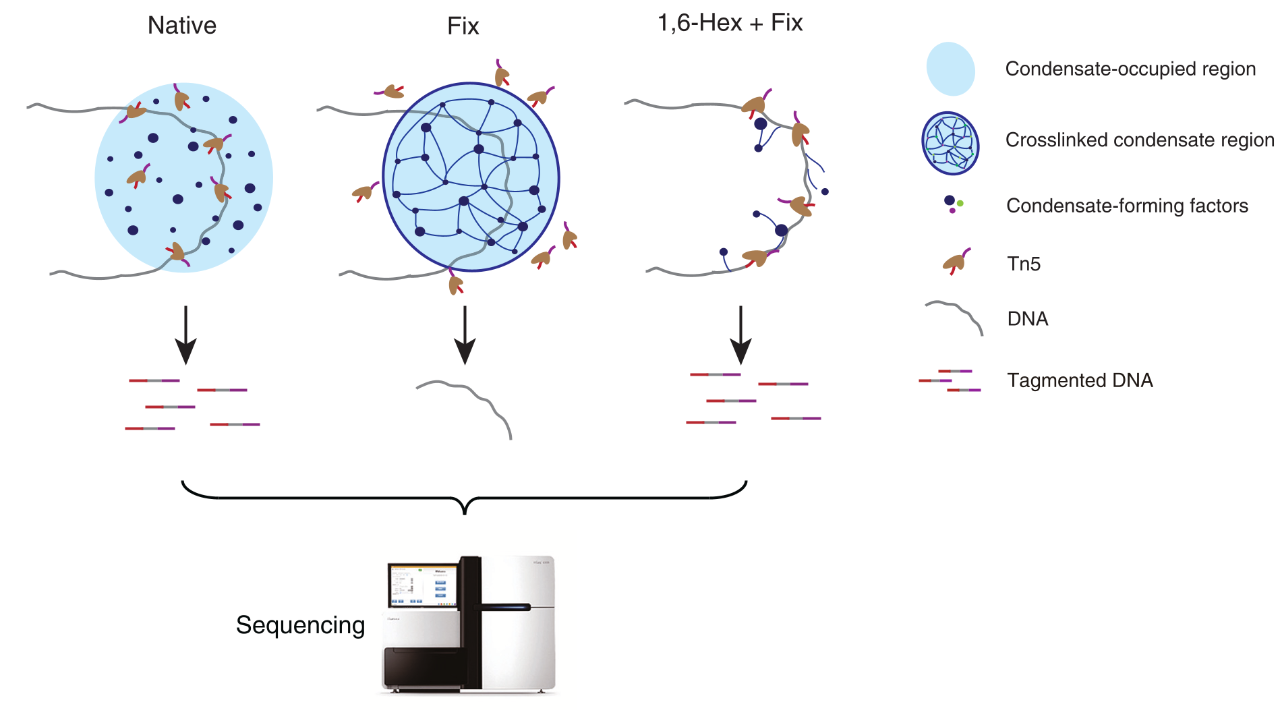
Workflow
ACC-seq utilizes the Tn5 transposase to assess differences in DNA fragment accessibility under various condensate-modulating conditions to identify condensate-occupied regions. The protocol involves three core treatment steps to modulate the state of chromatin condensates:
1. Native: No treatment is applied, maintaining the natural state of intracellular condensates.
2. Fix: Cells are fixed with formaldehyde, crosslinking condensates to their bound DNA.
3. 1,6-Hex+Fix: Cells are first treated with 1,6-hexanediol to disrupt condensate structures and release the bound DNA, followed by fixation.
Samples from these three conditions are subjected to Tn5 transposase cleavage. Since condensates sterically hinder Tn5 from cutting DNA, condensate-occupied regions exhibit reduced accessibility in the Fix sample but restored accessibility in the 1,6-Hex+Fix sample. The cleaved DNA fragments are then sequenced via high-throughput sequencing.
Applications
1. Genome-Wide Identification:
Mapping the occupancy regions of chromatin-bound condensates across the entire genome.
2. Gene Expression Regulation:
Investigating the impact of chromatin-bound condensates on gene expression.
3. TF Characterization:
Integrated analysis with other sequencing and imaging technologies to study the characteristics and functions of condensate-forming transcription factors (TFs).
4. Disease Mechanisms:
Analyzing the effects of disease-associated TF variants on condensate formation and transcriptional regulation.
5. Novel Discovery:
Discovery and characterization of new chromatin-bound condensates.
Advantages
1. High Sensitivity:
Capable of identifying condensates formed by direct DNA-binding proteins and capturing subtle regulatory changes. It can detect small condensates formed by a low number of TF molecules.
2. Robust Specificity:
Effectively distinguishes true condensate occupancy from false positives by analyzing differential accessibility patterns under specific modulation conditions.
3. Streamlined Workflow:
The experimental process is relatively simple, easy to operate, and requires a low cell input.
4. Comprehensive Insight:
Provides information on binding sites, expression levels, and functions of condensate-forming TFs.
Sample Requirements
Preparation: Cells must be processed according to the EPIBIOTEK® "One-step ACC-seq Pre-treatment Protocol."
1. Native Group: 5×10⁴ cells
2. Fix Group: 5×10⁴ cells
3. 1,6-Hex+Fix Group: 5×10⁴ cells
Live Cells
≥1.5×10⁵ cells/sample
Bioinformatic Analysis
Basic Analysis
1. Raw data processing (Adapter removal, QC).
2. Reference Genome Mapping.
3. Enrichment Region Identification (Peak Calling).
4. Identification of Hex-released open regions.
5. Peak Annotation.
6. GO Enrichment Analysis of Peak-associated genes.
7. KEGG Pathway Analysis of Peak-associated genes.
8. Motif Analysis of enriched regions.
9. Differential Peak analysis between groups (Screening, GO clustering, KEGG).
Advanced Analysis
1. Dual-action TFs prediction.
2. Combined TSS enrichment heatmaps.
3. GO/KEGG Chord plots.
4. IGV Visualization tracks (for 5 specific genes).
Demo
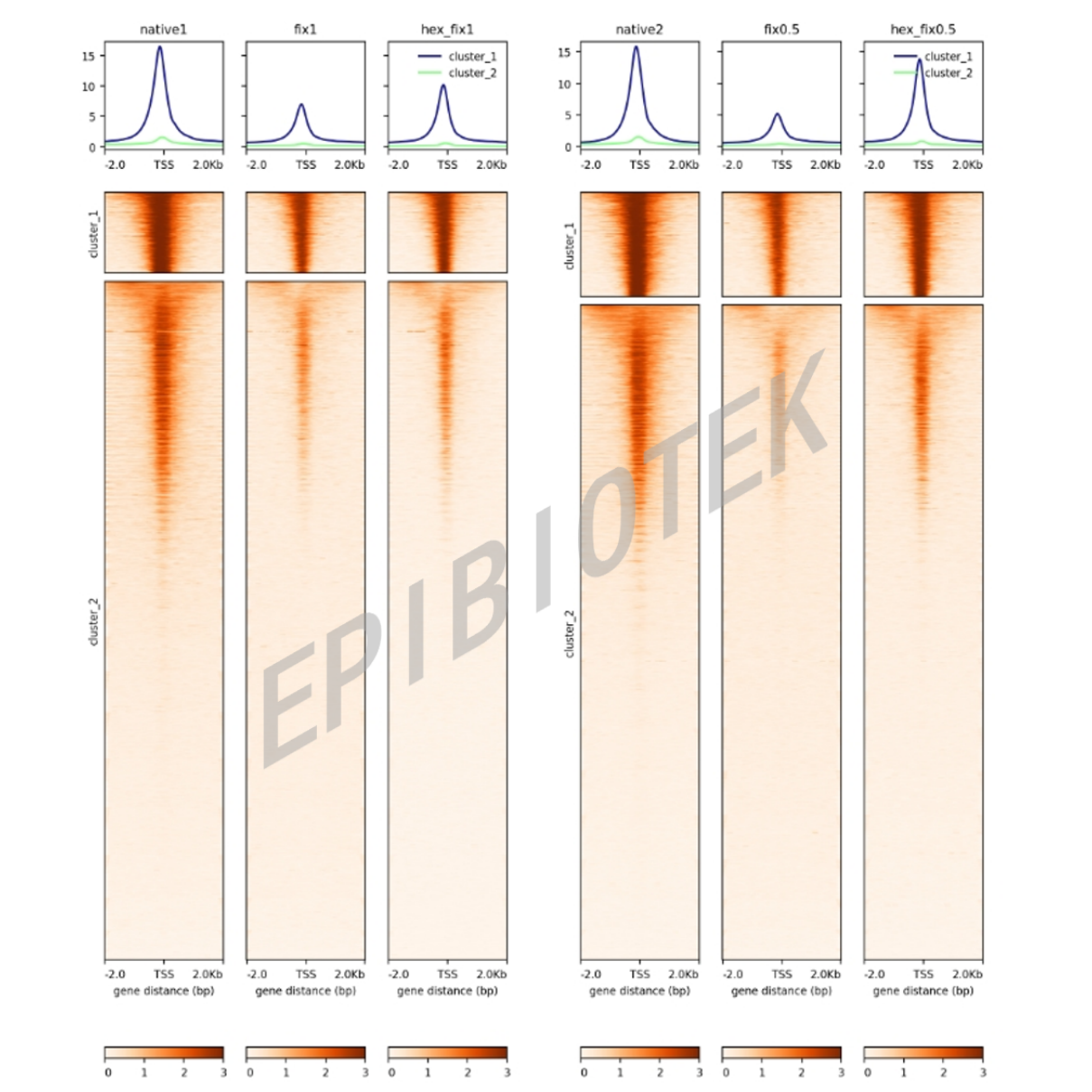
ACC-seq signal enrichment heatmap demonstrating a U-shaped structure. Cluster 1 represents "Consistent" open regions; Cluster 2 represents "Hex-released" open regions, which bind Dual-action TFs and affect chromatin stability.

Motif Analysis: Prediction of Dual-action TFs.
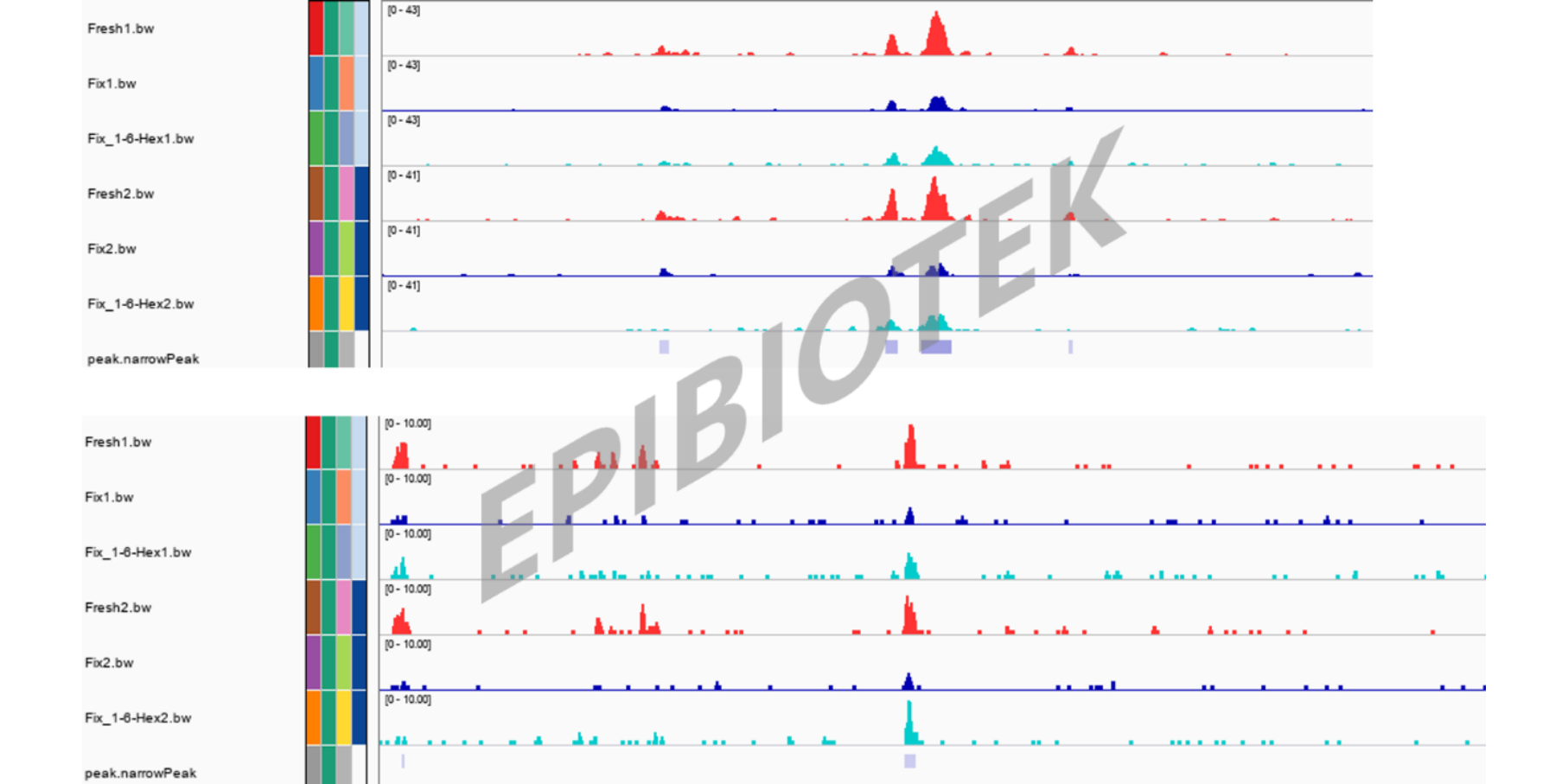
IGV Peak Visualization.
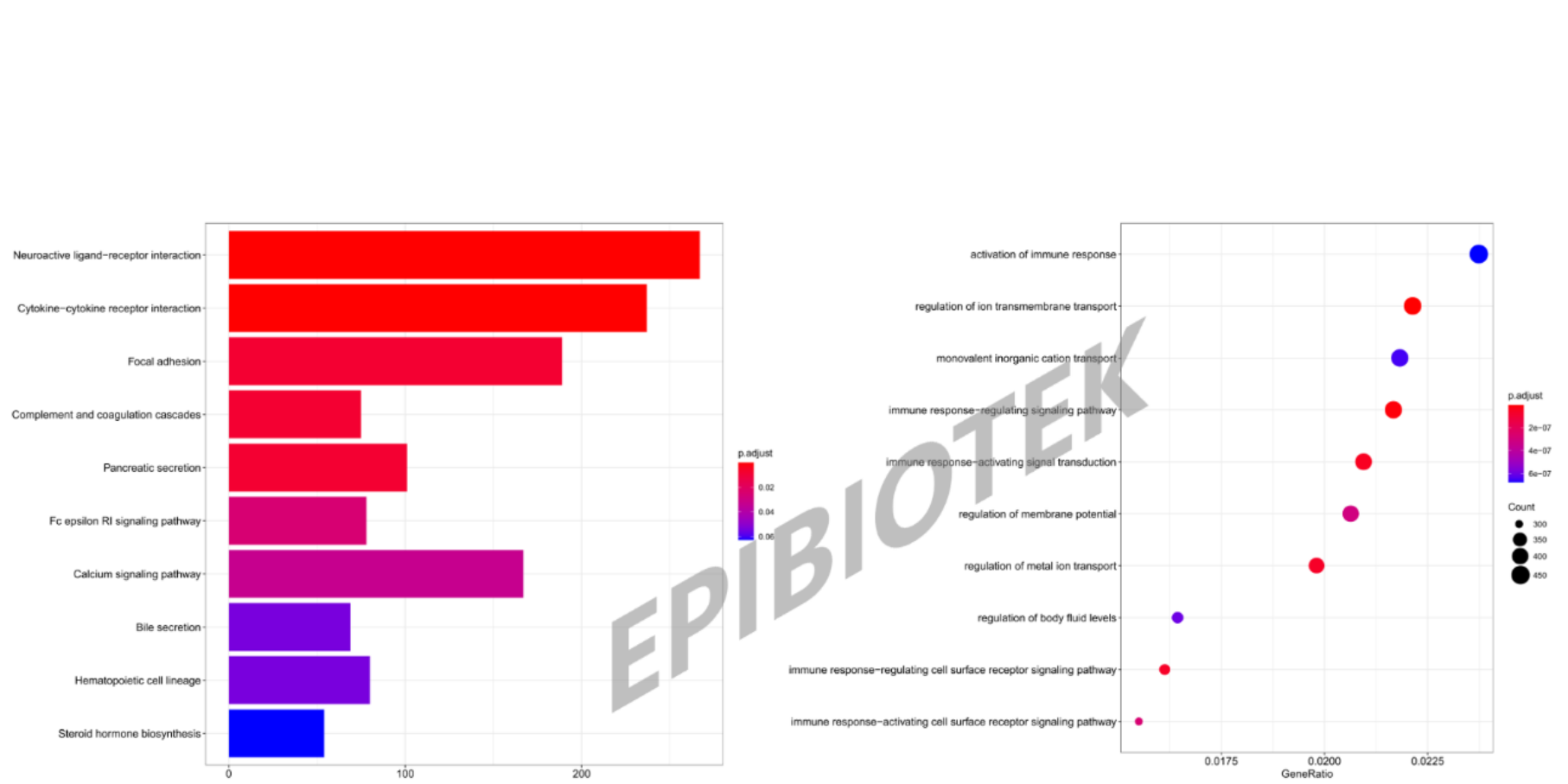
Bar charts for KEGG pathway analysis and bubble charts for GO enrichment analysis.
