HiChIP: Chromatin Conformation Sequencing Technology
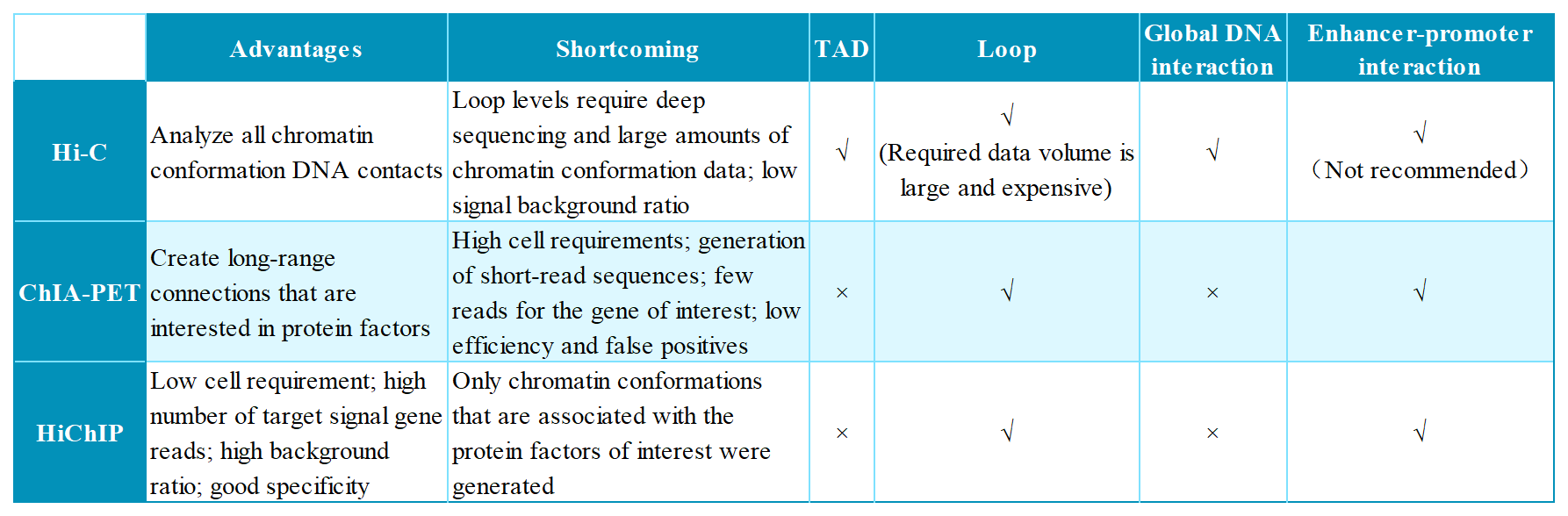
Table 1. Comparison of Hi-C, ChIA-PET and HiChIP techniques
Introduction
In situ Hi-C followed by chromatin immunoprecipitation (HiChIP) is a novel technique for analyzing chromatin conformation. It was developed by the laboratory of Howard Chang and published in Nature Methods in November 2016¹. HiChIP combines the features of Hi-C and ChIA-PET, further integrating them with transposase-mediated library construction techniques to obtain higher-resolution 3D chromatin structure information with reduced data requirements.
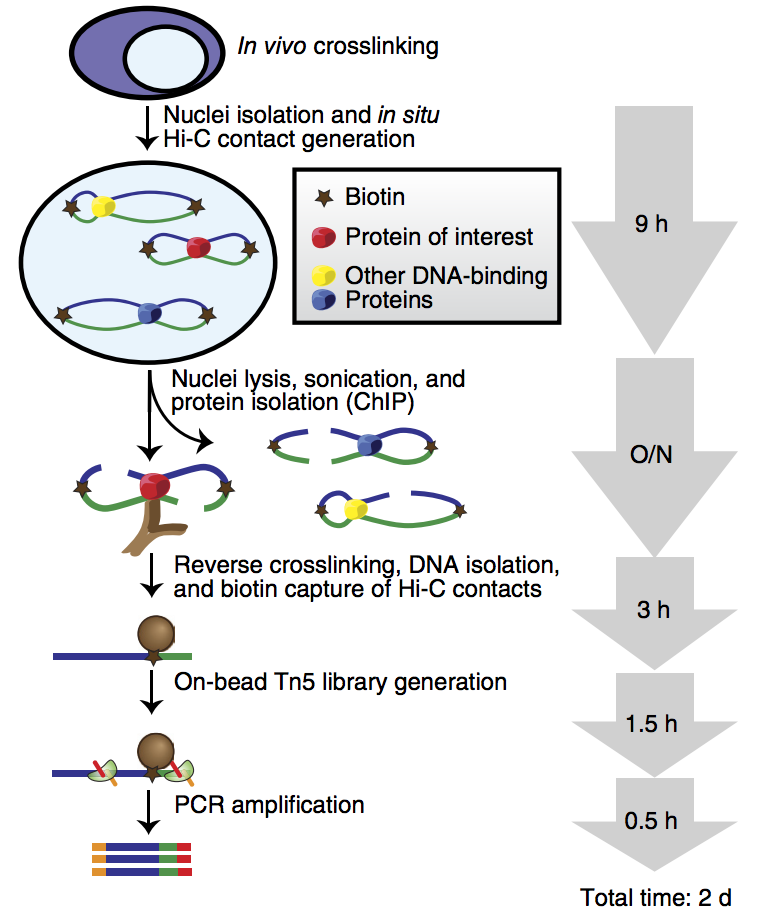
Figure 1. Workflow of HiChIP¹
Technical principle
HiChIP begins with in-nucleus cross-linking of intact cells, minimizing spurious contacts and maximizing capture efficiency of true chromatin interactions. Nuclei are isolated and subjected to in-situ Hi-C ligation; DNA ends are biotinylated before nuclear lysis and sonication. Chromatin immunoprecipitation (ChIP) is then performed with a locus-specific antibody, yielding DNA–protein complexes. After elution and reversal of cross-links, biotin pull-down enriches for Hi-C ligations, followed by library preparation and high-throughput sequencing.
Applications and Advantages
Application
1. Targeted analysis of chromatin-interaction conformations bound to a specific protein.
2. Elucidation of transcription factor mechanisms of action.
3. Investigation of how epigenetic modifications regulate gene expression.
Advantages
1. Delivers >10-fold higher yield of conformational information.
2. Requires >100-fold less starting material compared with ChIA-PET.
3. Needs fewer cells than Hi-C while providing a markedly higher signal-to-noise ratio.
4. Enables targeted acquisition of specific three-dimensional chromatin-structural information.
Sample Requirements
1. Prior preparation requires fixed cells, which should be provided as cell pellets and shipped on dry ice. Please consult technical support for cell fixation protocols.
2. If prior cell fixation is not feasible, please provide live cells with the appropriate culture medium, and ship at ambient temperature.
Live cells
Types
≥1×10⁷ crosslinked cells/sample
Cell Count
Limited to human, mouse, and rat cells and tissues
Species
Bioinformatic Analysis
1. Sequence alignment.
2. HiChIP experimental quality control analysis (alignment and valid interaction pairs).
3. HiChIP interaction heatmaps (multiple resolutions).
4. Loops identification.
5. Loops annotation (enhancer-promoter interactions).
6. Hub super-enhancer identification.
7. Generation of visualization files.
8. Differential analysis of loops.
9. GO and KEGG analysis.
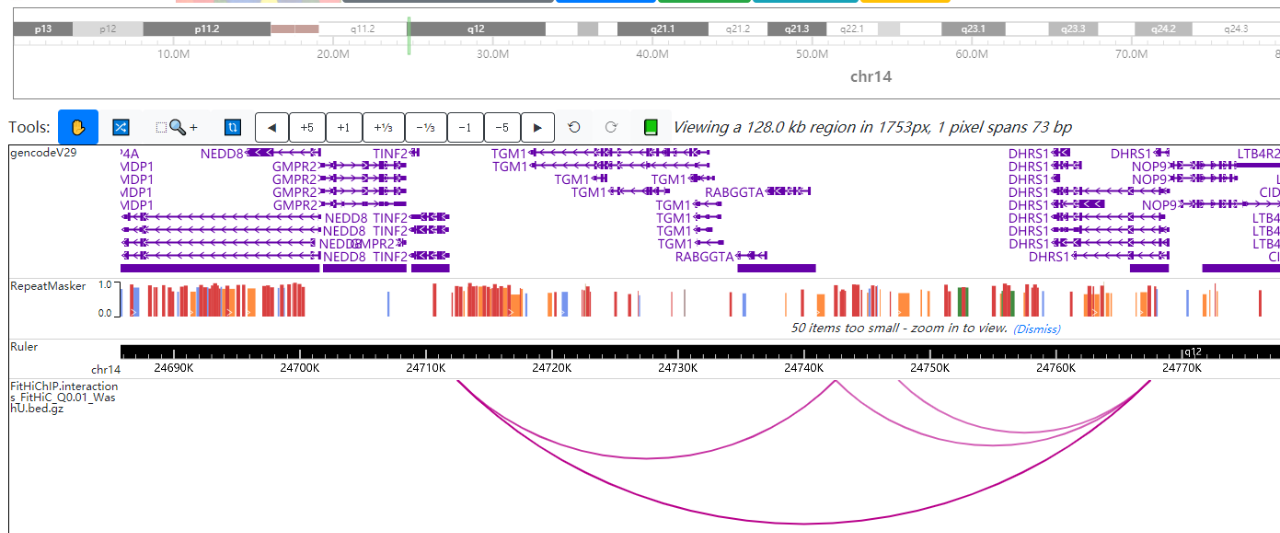
Figure 2. Visualization Loop
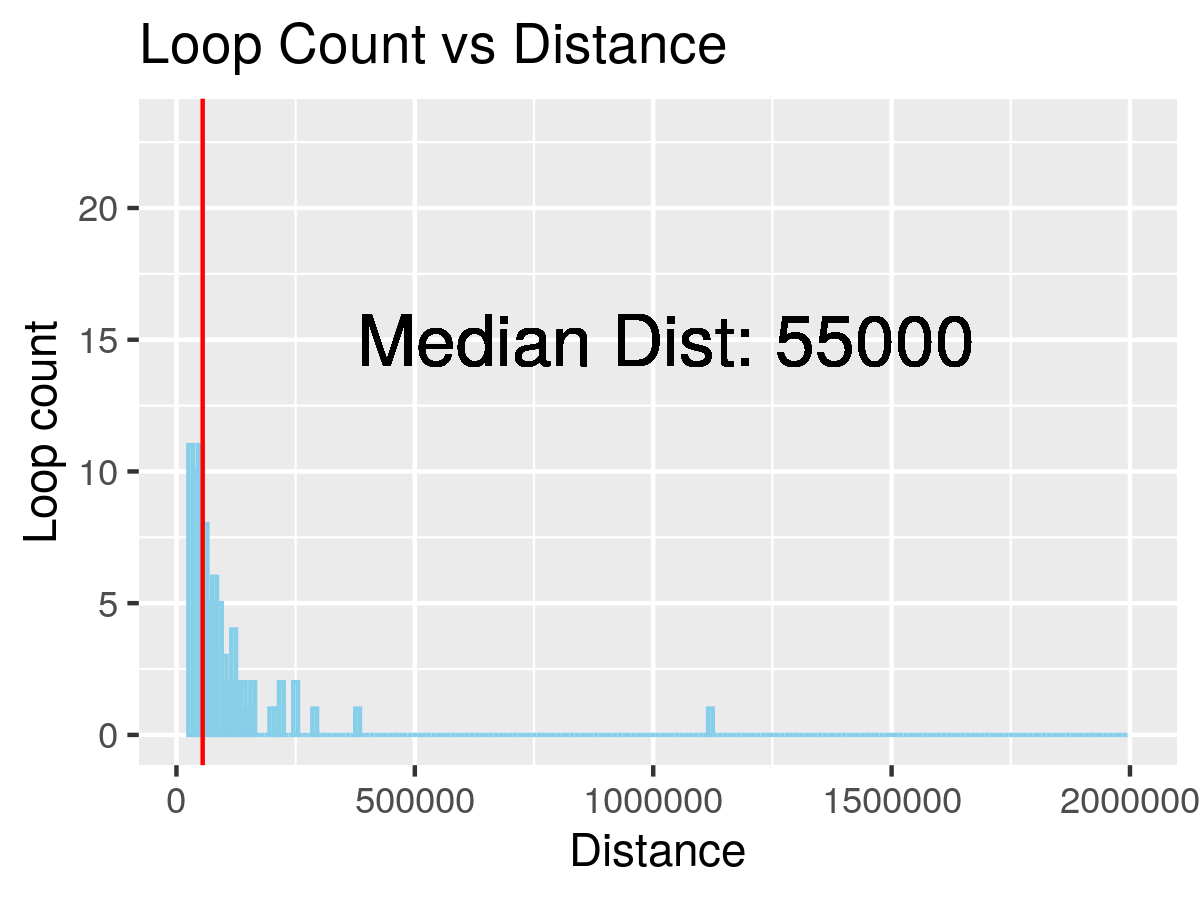
Figure 3. Loop distance histogram
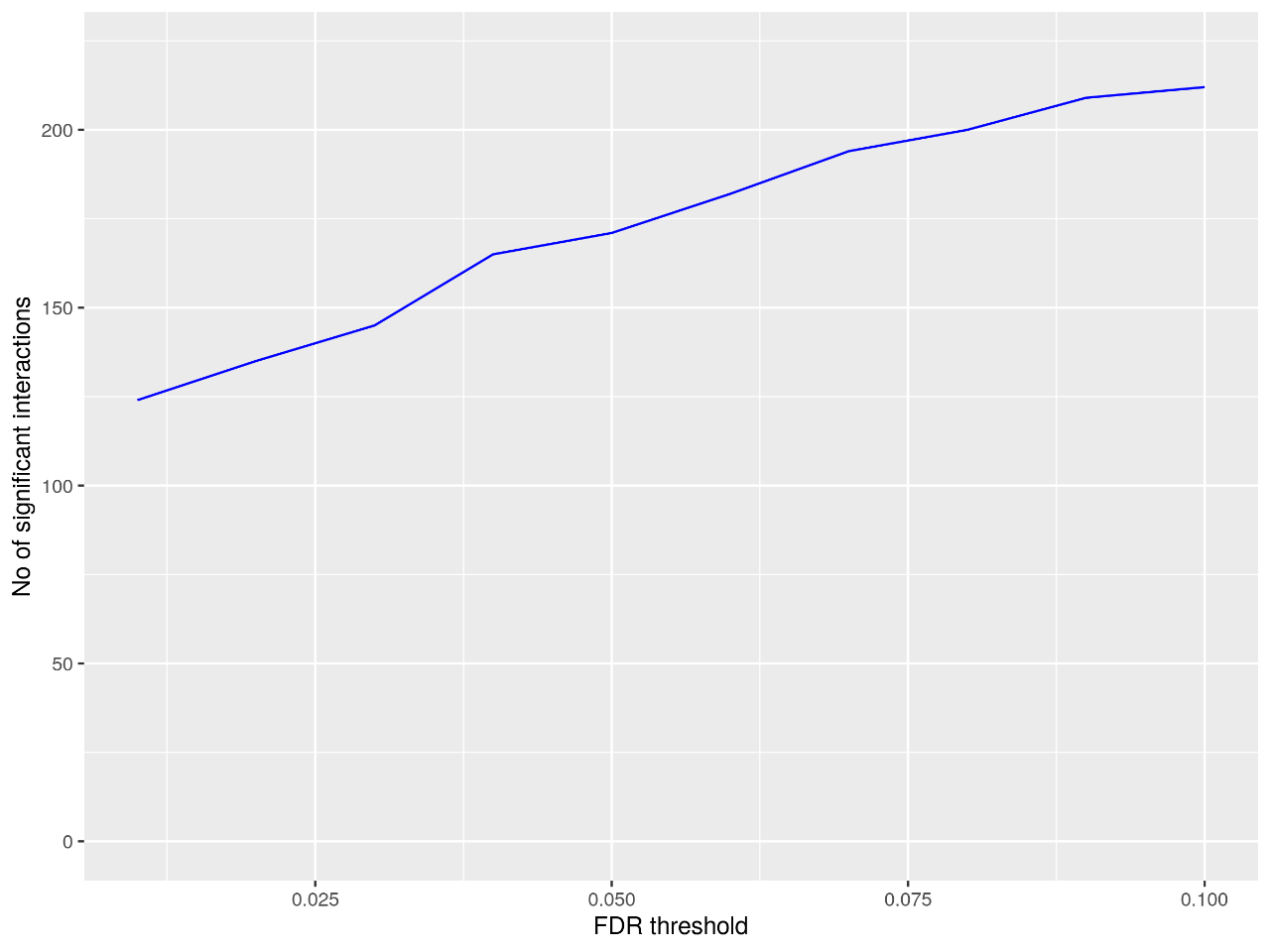
Figure 4. Statistical diagram of significant interactions
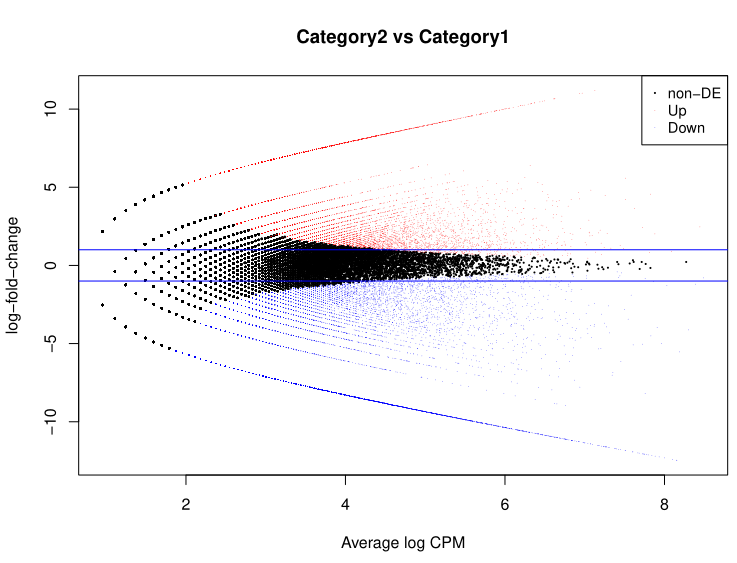
Figure 5. Loop MA Figure
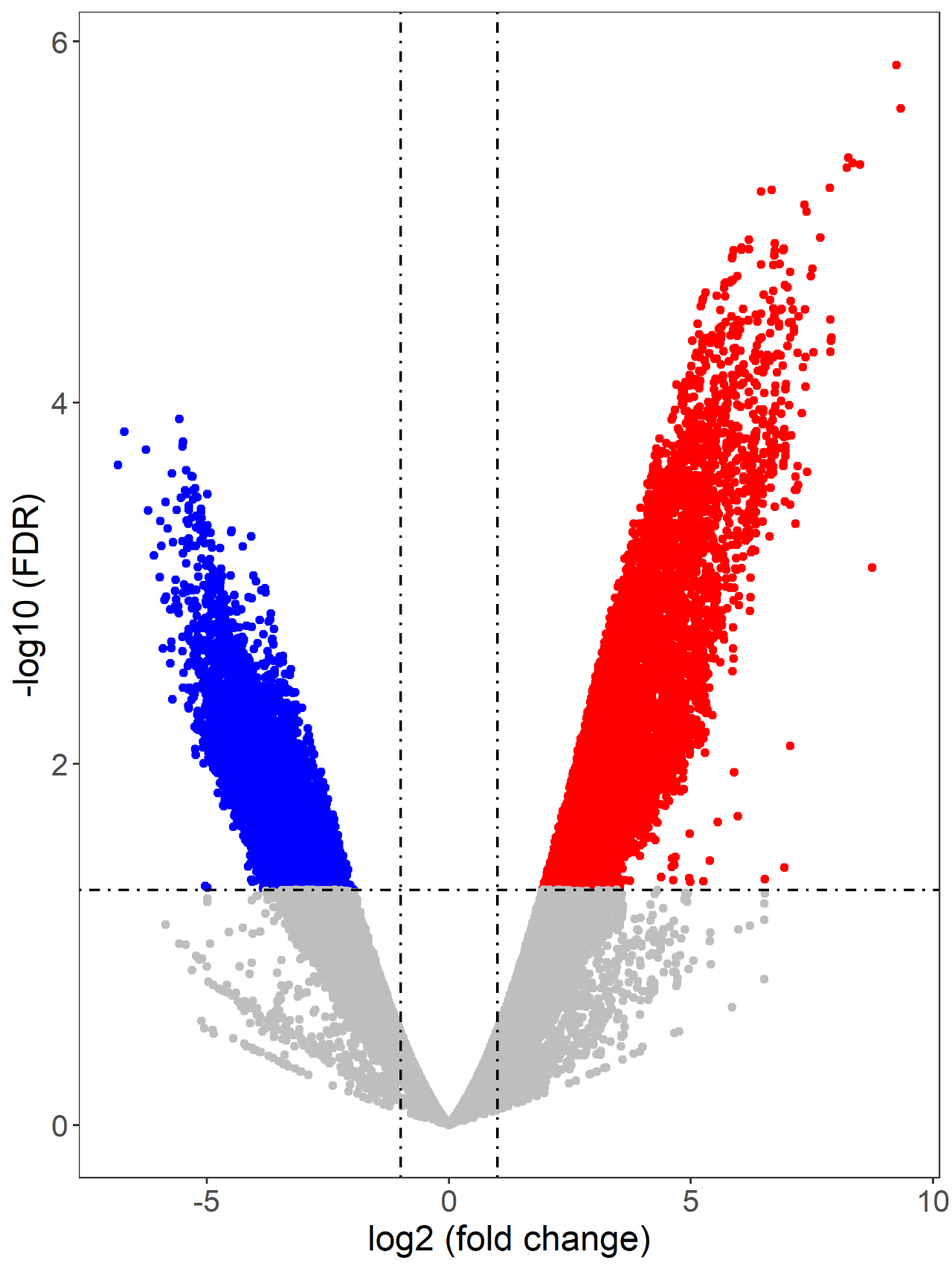
Figure 6. Loop volcano diagram
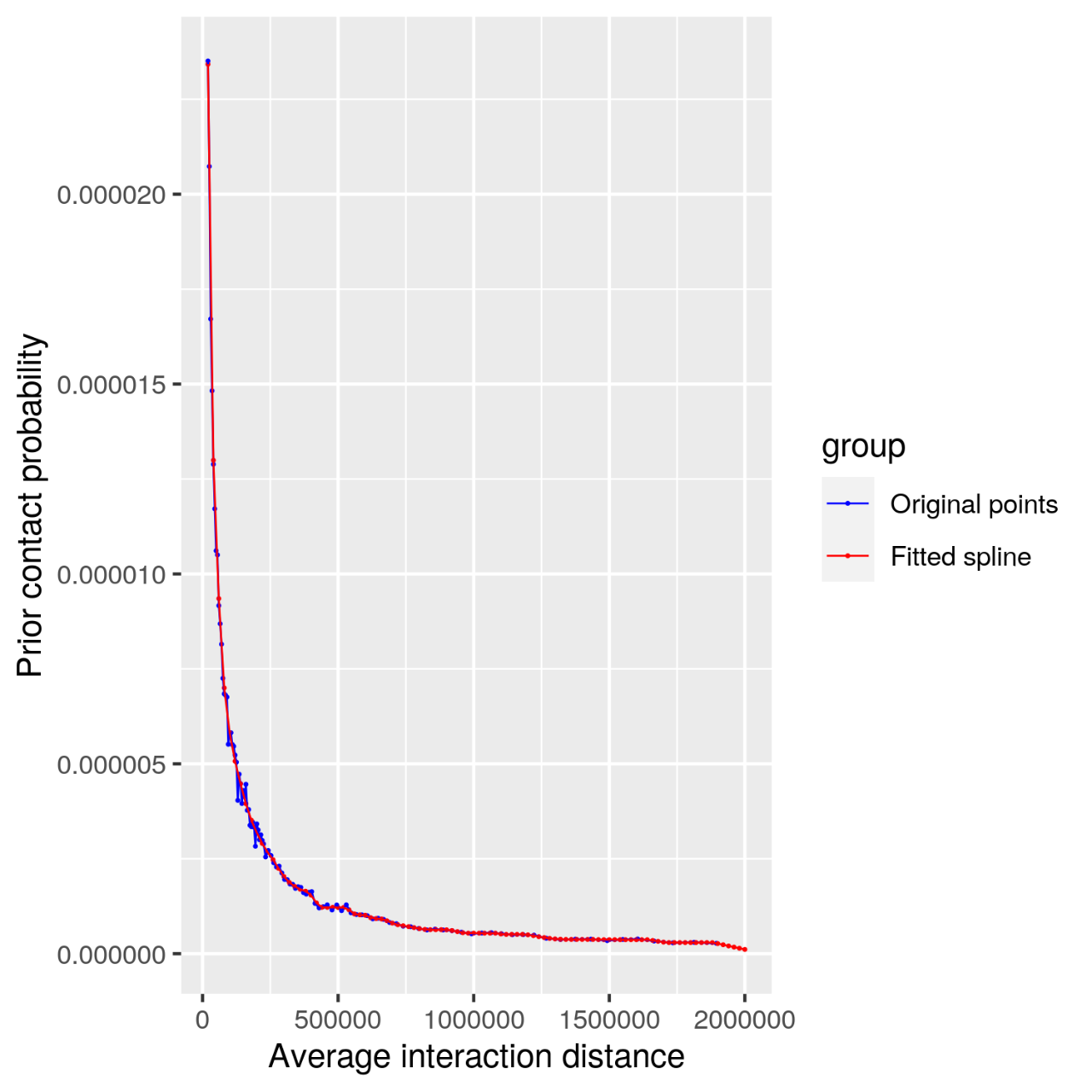
Figure 7. Contact filter diagram
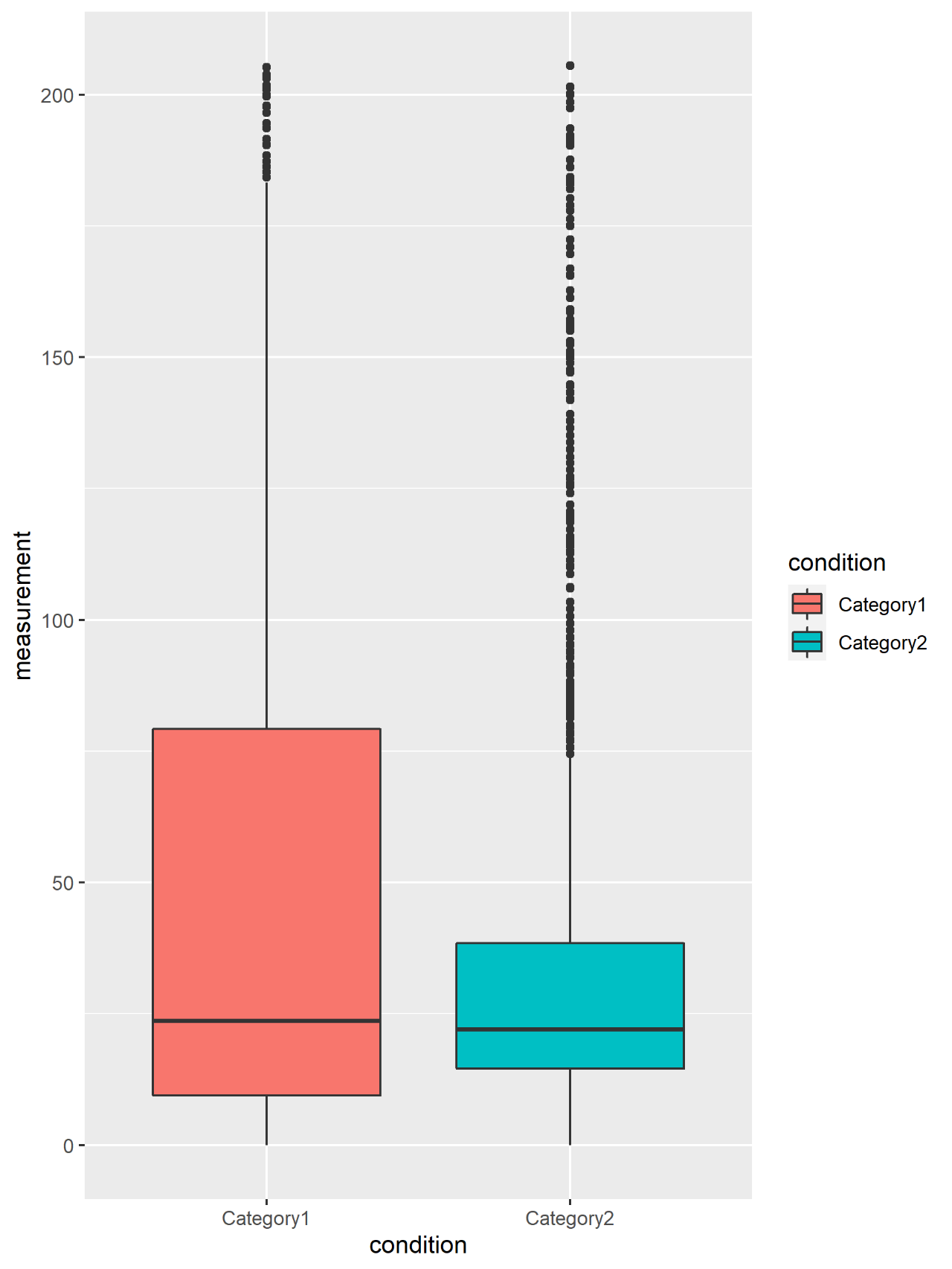
Figure 8. Loop box diagram
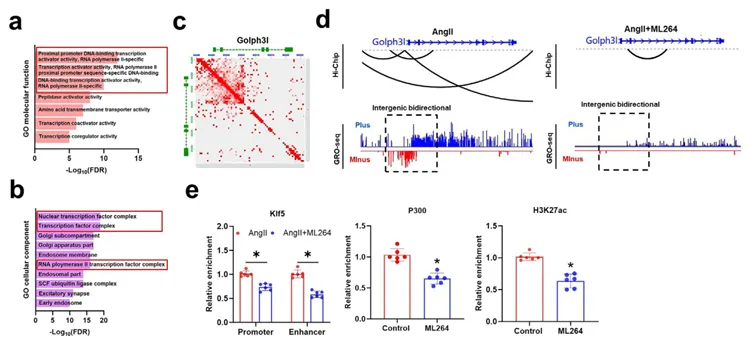
Figure 9. Combined analysis of HiChIP and GRO‑seq
Project Article
《ADVANCED SCIENCE》 Ythdc1-p300-Klf5 Complex-Mediated Golgi Dysfunction Promotes Aortic Aneurysm(2)
This study elucidates the molecular mechanism by which apoptosis-induced proliferation in vascular smooth muscle cells promotes the development of aortic aneurysm (AD/AAA). The study found that AngII stimulation increases the m6A modification level of lncRNA Gm40097, which recruits the reader protein Ythdc1 and facilitates its formation of a complex with p300 and the transcription factor Klf5, thereby remodeling chromatin states to activate GOLPH3L transcription. High expression of GOLPH3L induces Golgi apparatus condensation, promotes the secretion of inflammatory factors such as TNF-α, and ultimately drives disease progression. Additionally, the study constructed a postoperative mortality risk prediction model based on clinical features and functional SNPs.
References
[1] Mumbach MR, Rubin AJ, Flynn RA, et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat Methods. 2016;13(11):919-922.
[2] Wang WL, Song ZX, Bu SM, et al. Ythdc1-p300-Klf5 Complex-Mediated Golgi Dysfunction Promotes Aortic Aneurysm. Adv Sci (Weinh). Published online November 29, 2025. (EPIBIOTEK® Project Publications)
[3] Jia Q, Tan Y, Li Y, Wu Y, Wang J, Tang F. JUN-induced super-enhancer RNA forms R-loop to promote nasopharyngeal carcinoma metastasis. Cell Death Dis. 2023;14(7):459. Published 2023 Jul 21.(EPIBIOTEK® Project Publications)
[4] Wang W, Song F, Feng X, et al. Functional Interrogation of Enhancer Connectome Prioritizes Candidate Target Genes at Ovarian Cancer Susceptibility Loci. Front Genet. 2021;12:646179. Published 2021 Mar 19.(EPIBIOTEK® Project Publications)
