cfWGBS
cfWGBS
Whole-genome bisulfite sequencing of cell-free DNA
Liquid biopsy technology has demonstrated immense potential in disease diagnosis, therapeutic monitoring, and prognostic evaluation due to its non-invasive nature, convenience, and reproducibility. As a pivotal component of liquid biopsy, cell-free DNA (cfDNA)—fragmented DNA found in bodily fluids carrying genetic information from various sources like tumor or fetal cells—provides new perspectives for early disease diagnosis and precision medicine.
Whole Genome Bisulfite Sequencing (WGBS) is widely recognized as the "gold standard" for DNA methylation research [1-2]. By utilizing bisulfite treatment, WGBS generates a comprehensive, single-base resolution map of DNA methylation across the entire genome.
EPIBIOTEK® offers cfWGBS sequencing services (applying WGBS technology to cfDNA). This service provides a powerful tool for deeply analyzing disease mechanisms, discovering novel biomarkers, and guiding personalized treatment strategies.
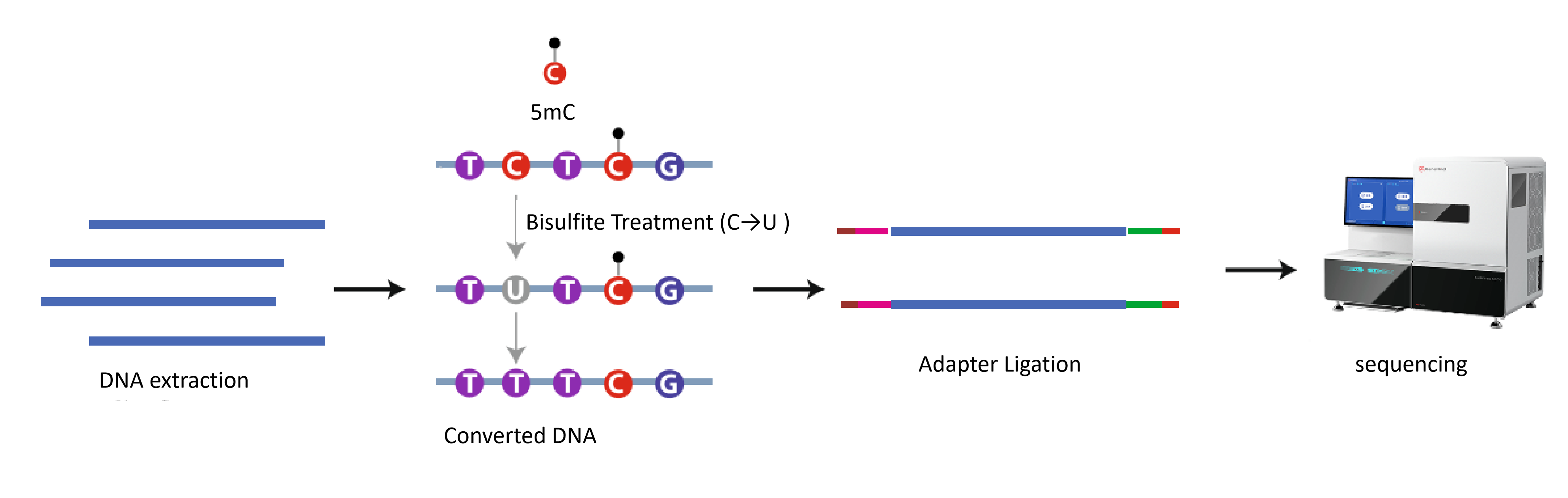
Workflow
1. DNA Extraction: Isolation of cfDNA from the target samples (e.g., plasma).
2. Bisulfite Conversion: Treatment of DNA with bisulfite. Unmethylated cytosines (C) are converted to uracils (U), while methylated cytosines (mC) remain unchanged.
3. Library Construction: Fragmentation of the converted DNA (if necessary) and ligation of sequencing adapters to construct the DNA library.
4. High-Throughput Sequencing: Sequencing of the quality-controlled library.
5. Bioinformatics Analysis: Alignment of reads to the reference genome and determination of the methylation level of each cytosine based on the C-to-T ratio.
Applications
1. Methylome Profiling: Construction of genome-wide, single-base resolution DNA methylation maps.
2. Physiological Mechanisms: Investigation of epigenetic mechanisms underlying physiological processes such as embryonic development and aging.
3. Biomarker Discovery: Identification of disease-associated methylation biomarkers, particularly for early cancer diagnosis.
Advantages
1. High Precision: Identifies CpG sites with single-base resolution.
2. Comprehensive Coverage: As the gold standard, it covers the entire genome, providing the most complete methylation information available.
3. Non-Invasive Sampling: Unlike traditional tissue biopsies, cfWGBS requires only standard blood samples.
Sample Requirements
Plasma/Serum
1–4 mL/sample
cfDNA
≥15 ng
Bioinformatics Analysis
1. Data Quality Control & Alignment
a. Genomic mapping statistics.
2. Global Methylation Assessment
a. Average methylation levels.
b. Genome-wide methylation distribution.
3. Methylation Site Statistics & Comparative Analysis
a. Correlation analysis of methylation levels.
b. PCA Analysis.
c. Cluster analysis of methylation levels.
4. Differentially Methylated Analysis
a. Annotation of Differentially Methylated Regions (DMRs).
b. Length distribution of DMRs.
c. Methylation level distribution within DMRs.
d. Clustering heatmaps of DMRs.
e. Visualization of DMRs.
f. GO and KEGG analysis for genes associated with DMRs.
Demo
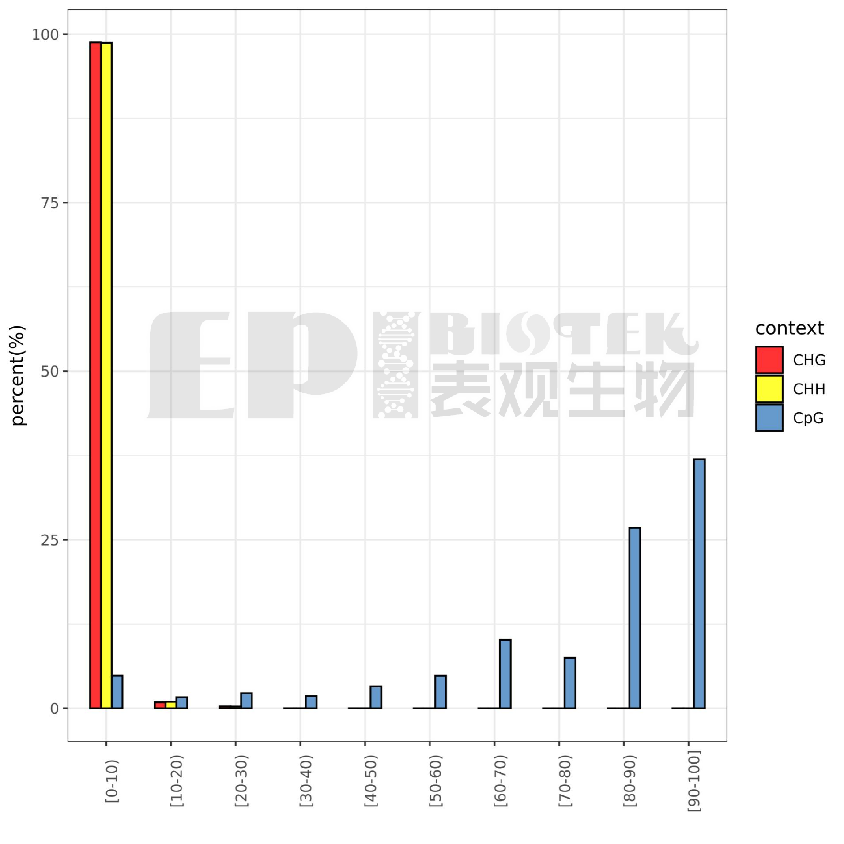
Distribution of methylation levels at specific sites
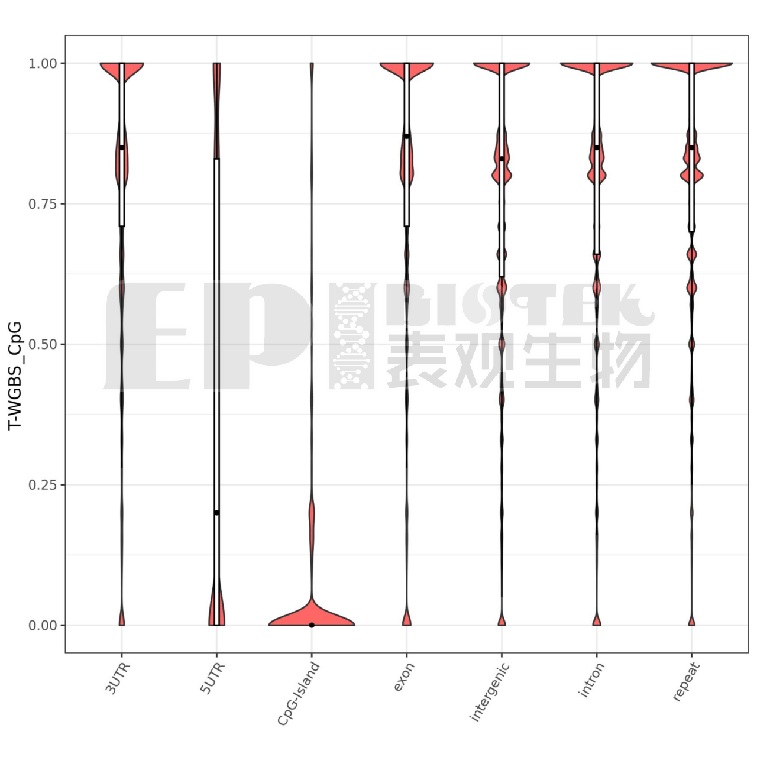
Methylation profiles across genomic elements
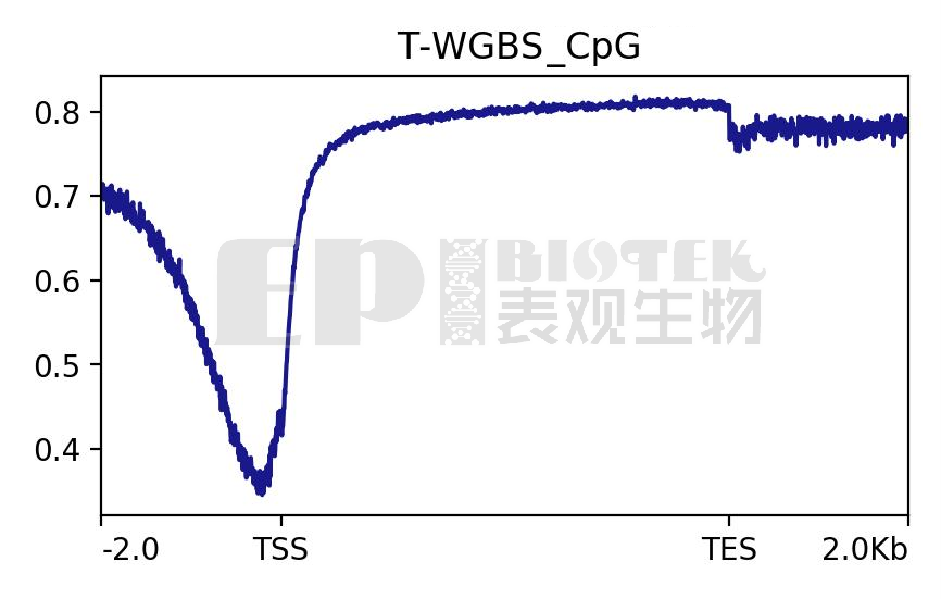
Average methylation profile around TSS
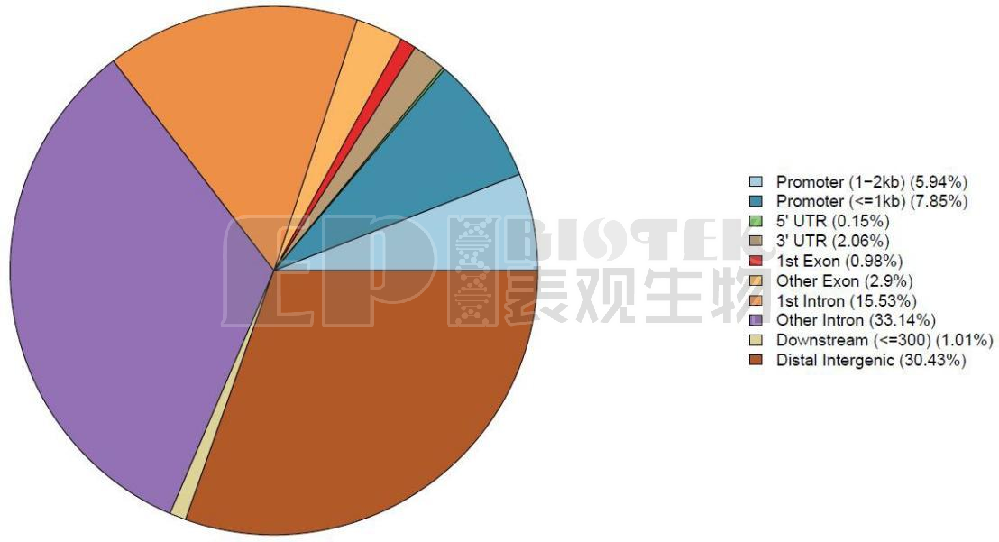
Distribution of DMR types
References
[1]Guo H, Zhu P, Yan L, et al. The DNA methylation landscape of human early embryos. Nature.2014.
[2] Ehsan Habibi, et al. Whole-Genome Bisulfite Sequencing of Two Distinct Interconvertible DNA Methylomes of Mouse Embryonic Stem Cells. Cell Stem Cell (2013), 13(3), 360-369.
[3] Masser DR, Hadad N, Porter H, et al. Analysis of DNA modifications in aging research. Geroscience. 2018;40(1):11-29.
