cfDNA fragmentomics
cfDNA fragmentomics
Cell-free DNA (cfDNA) refers to nucleic acid fragments released into the extracellular space through apoptosis, necrosis, or active secretion by cells, including cancer cells. cfDNA carries valuable molecular information such as mutations, copy number variations (CNV), methylation patterns, and tissue-of-origin signatures (1).
cfDNA fragmentomics represents a cutting-edge liquid biopsy approach for non-invasive cancer detection and medical research. It characterizes the distinctive fragmentation patterns of tumor-derived DNA compared to DNA from other tissues, and has gained increasing attention in recent years (2).
this service utilizes whole-genome sequencing (WGS) to analyze fragmentation patterns in blood-derived cfDNA. Key features include Fragment Size Ratio (FSR), Fragment Size Distribution (FSD), End Motif (EDM), Breakpoint Motif (BPM), CNV, SNP, and nucleosome footprints, which reflect chromatin accessibility and gene expression states. These characteristics are closely linked to genomic organization, cell death mechanisms, and epigenetic modifications (3). By distinguishing differences in cfDNA fragmentation patterns and studying their associations with DNA methylation, histone modifications, and chromatin structure, the service infers cancer-related epigenetic alterations. It is suitable for large-scale population screening studies.
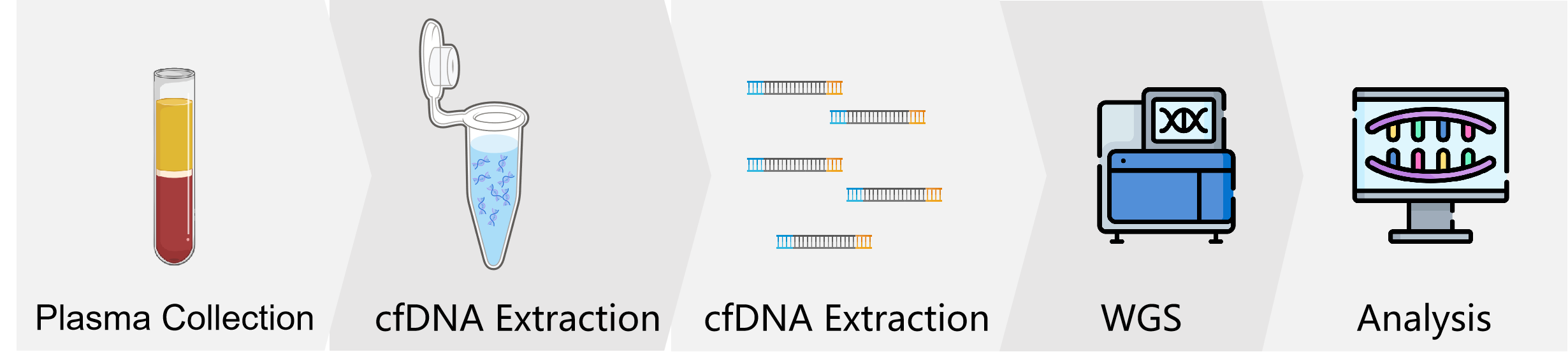
Workflow
Applications
1. Multi-cancer early detection and diagnosis
2. Tumor tissue-of-origin identification
3. Transplant rejection monitoring
4. Non-invasive prenatal testing
5. Molecular prognostic biomarker discovery
Sample Requirements
1. Type: Plasma, 2 mL per sample
2. Species: Human only
Bioinformatic Analysis
1. Raw data quality control
2. Genome alignment
3. FSR calculation
4. FSD analysis
5. EDM and BPM profiling
6. CNV and SNP variant detection
7. Nucleosome footprint analysis
8. Single- and multi-cancer prediction modeling
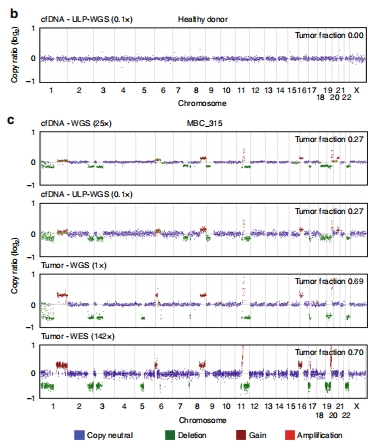
FSD (4)
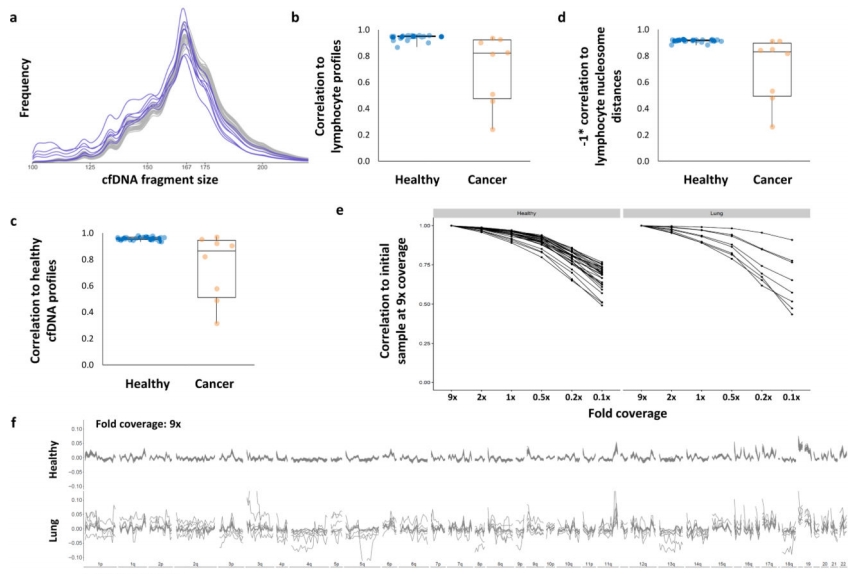
FSR statistics comparing healthy individuals and cancer patients (5)
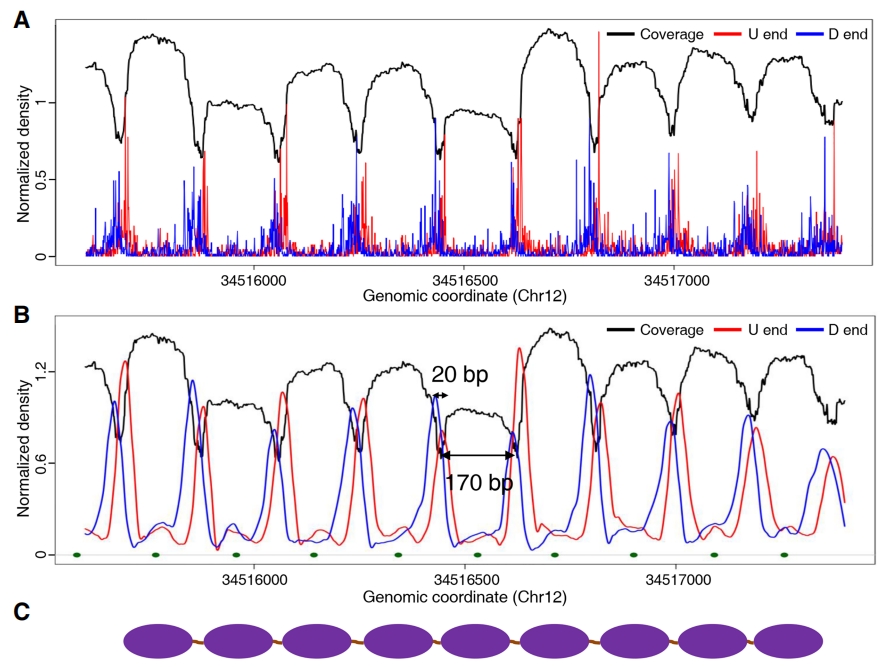
Nucleosome distribution patterns (6)
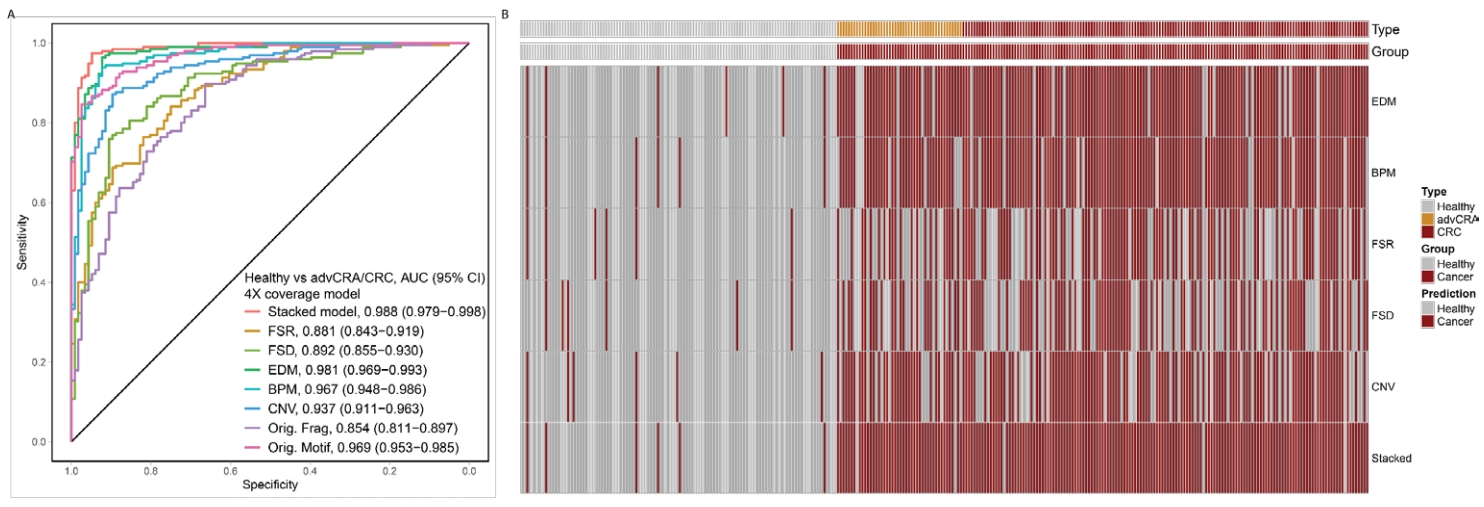
Multi-cancer prediction model AUC and feature heatmap (7)
References
[1] Wan JCM, Massie C, Garcia-Corbacho J, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017;17(4):223-238.
[2] Tsui WHA, Jiang P, Lo YMD. Cell-free DNA fragmentomics in cancer. Cancer Cell. 2025;43(10):1792-1814.
[3] Ding SC, Lo YMD. Cell-Free DNA Fragmentomics in Liquid Biopsy. Diagnostics (Basel). 2022;12(4):978. Published 2022 Apr 13.
[4] Adalsteinsson VA, Ha G, Freeman SS, et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat Commun. 2017;8(1):1324. Published 2017 Nov 6.
[5] Cristiano S, Leal A, Phallen J, et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature. 2019;570(7761):385-389.
[6] Sun K, Jiang P, Cheng SH, et al. Orientation-aware plasma cell-free DNA fragmentation analysis in open chromatin regions informs tissue of origin. Genome Res. 2019;29(3):418-427.
[7] Wang S, Meng F, Li M, et al. Multidimensional Cell-Free DNA Fragmentomic Assay for Detection of Early-Stage Lung Cancer. Am J Respir Crit Care Med. 2023;207(9):1203-1213.
