cfChIP-seq
Product Introduction
cell-free DNA chromatin immunoprecipitation sequencing
cell-free DNA chromatin immunoprecipitation sequencing (cfChIP-seq) is an innovative liquid-biopsy technology that opens a new window for decoding the gene-regulatory landscape of tumor cells by profiling histone modifications on circulating cfDNA. By employing highly specific antibodies, the method captures free nucleosomes from minute volumes of plasma and couples this enrichment with high-throughput sequencing to achieve quantitative mapping of diverse histone marks on cfDNA. Consequently, cfChIP-seq provides direct readouts of transcriptional activity, cellular state, and expression levels of key genes in malignant cells, complementing the limitations of conventional cfDNA analyses. Owing to its exceptional sensitivity and rich information content, the technology is poised to play a pivotal role in early cancer detection, personalized therapy selection, and real-time treatment-response monitoring, thereby advancing the clinical application of liquid biopsy in precision medicine.
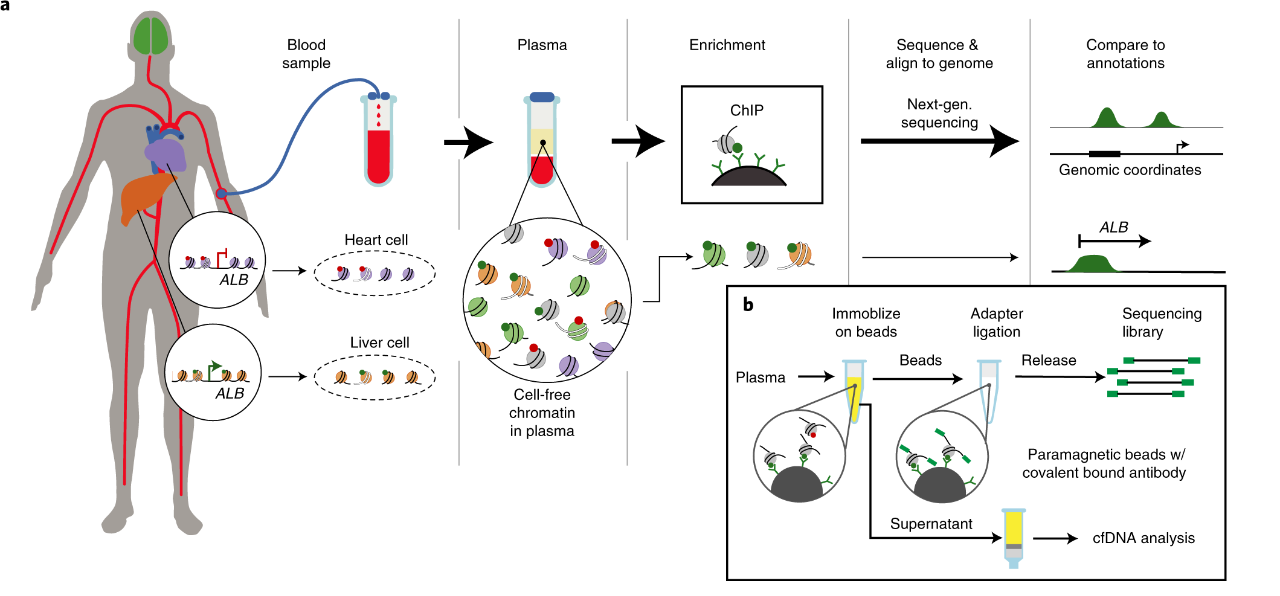
Figure 1. Workflow of cfChIP-seq(1)
Advantages
1. Only 1mL of plasma is required for analysis
2. Acquire information at multiple levels simultaneously
3. Can accurately quantify histone modifications on cfDNA
Application
1.The histone modification map of free nucleosomes in blood samples was analyzed
2.By analyzing histone modification patterns, different tumor subtypes can be accurately identified
3.It can reflect the dynamic response of tumor cells to treatment sensitively
4.Develop new liquid biopsy disease markers
5.The histone modification level of specific transcription factor binding sites was analyzed to find potential therapeutic targets
Sample Requirements
Types
Plasma/Serum, 1-4 mL/sample
Species
Limited to human, mouse, and rat cells; other species require evaluation.
Bioinformatic Analysis
1.Raw reads trimming and quality control
2.Alignment to the reference genome
3.Identification of enriched regions (Peak Calling)
4.Peaks Annotation
5.GO analysis of peak-associated genes
6.KEGG analysis of peak-associated genes
7.Motif analysis of enriched regions
8.Screening, GO functional cluster analysis, and KEGG analysis between groups of differential peak-associated genes
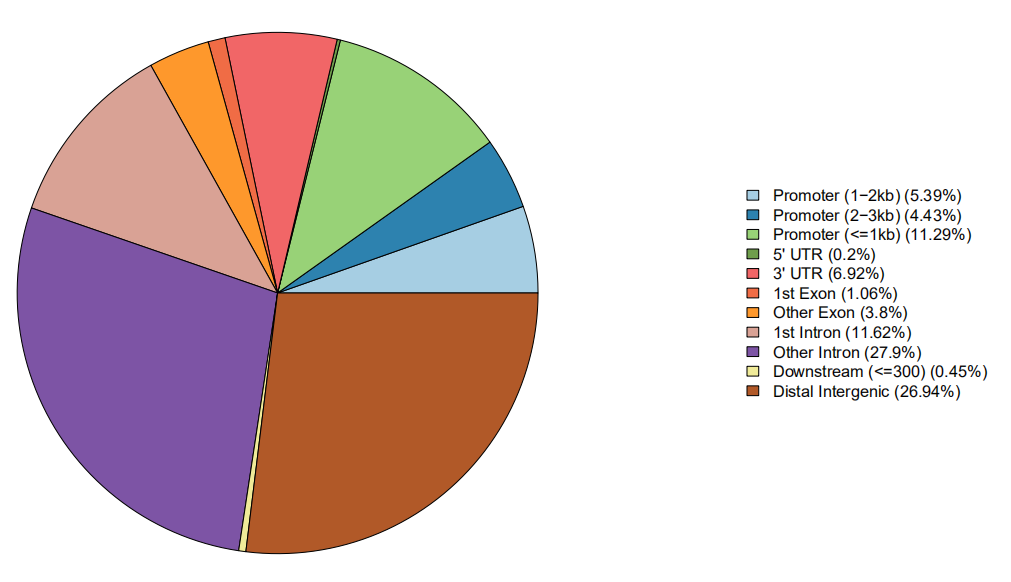
Figure 2. Genome Distribution Pie Chart
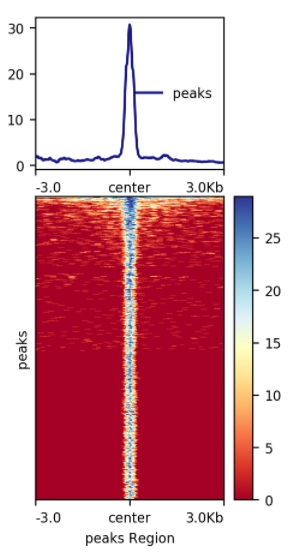
Figure 3. Signal Enrichment Plot and Distribution Heatmap
Related Articles
Nat Med:Liquid biopsy epigenomic profiling for cancer subtyping(2)
Researchers applied cfChIP-seq to 1,268 plasma samples collected from 433 patients with advanced-stage cancers and, by profiling the promoter- and enhancer-associated histone modifications H3K4me3 and H3K27ac, accurately discriminated among diverse malignancies—including neuroendocrine tumors, gastrointestinal carcinomas, and colorectal cancer. The circulating histone-mark patterns exhibited strong concordance with gene-expression levels in matched tumor tissues, faithfully recapitulating the epigenetic state of the underlying disease. The investigators further demonstrated that cfChIP-seq can detect epigenetic alterations linked to therapeutic resistance, such as activation of the androgen-receptor enhancer in prostate cancer, thereby offering valuable biomarkers for predicting treatment response and guiding the development of novel therapeutic strategies.
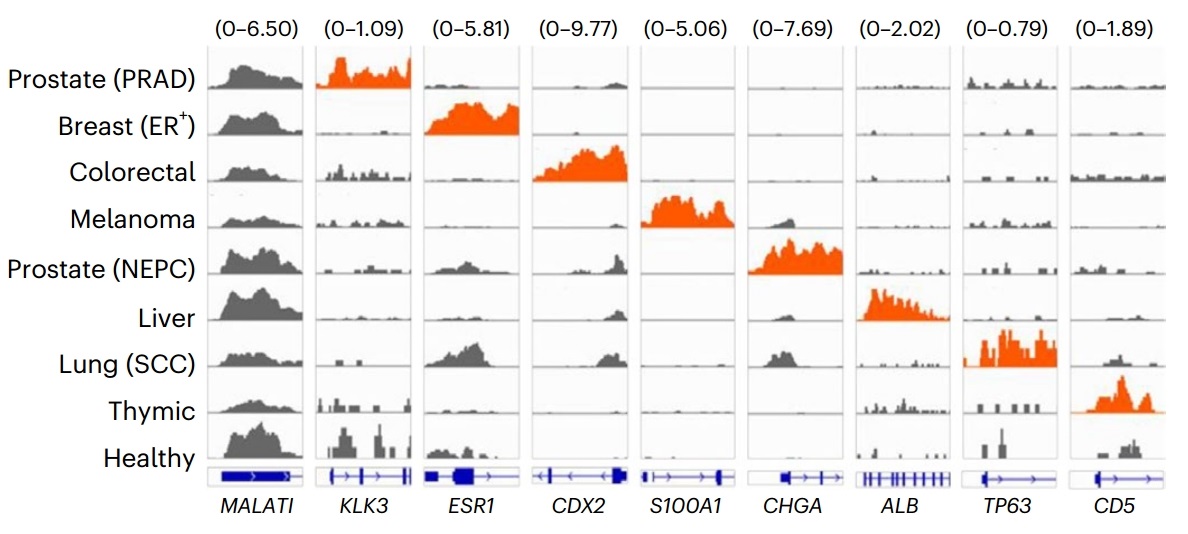
Figure 4. Detection of promoter activity of diagnostic marker genes in different cancer types in plasma samples by cfChIP-seq
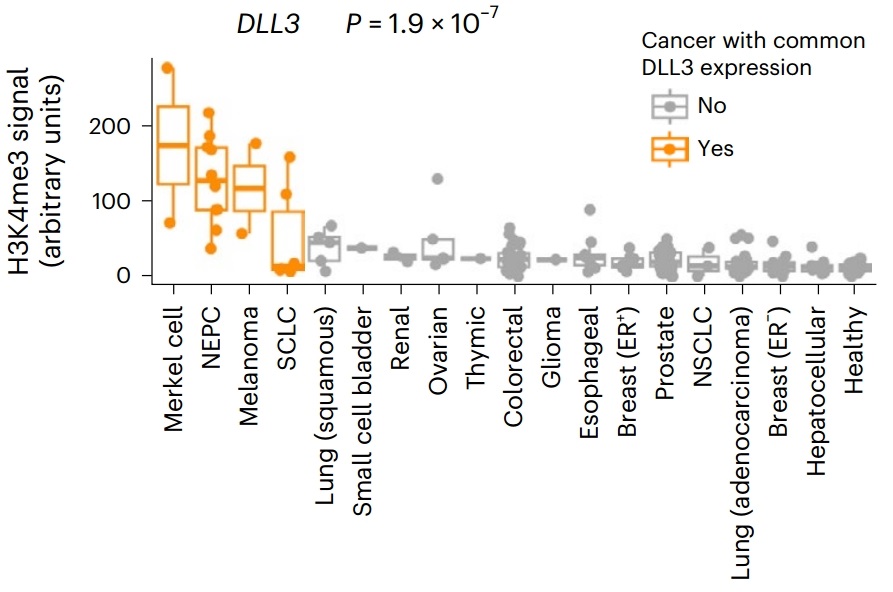
Figure 5. Distribution of H3K4me3 signals in the DLL3 gene promoter in different cancer types
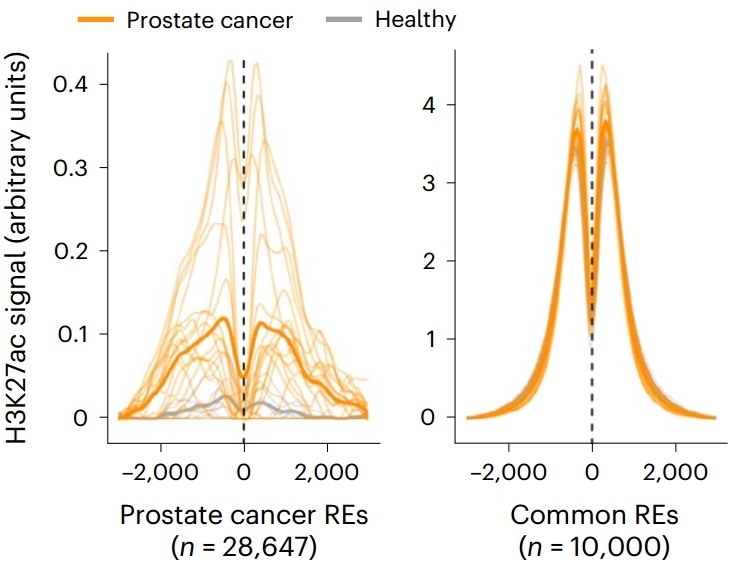
Figure 6. Enrichment of H3K27ac signal in tumor-specific enhancer regions in plasma samples of prostate cancer patients
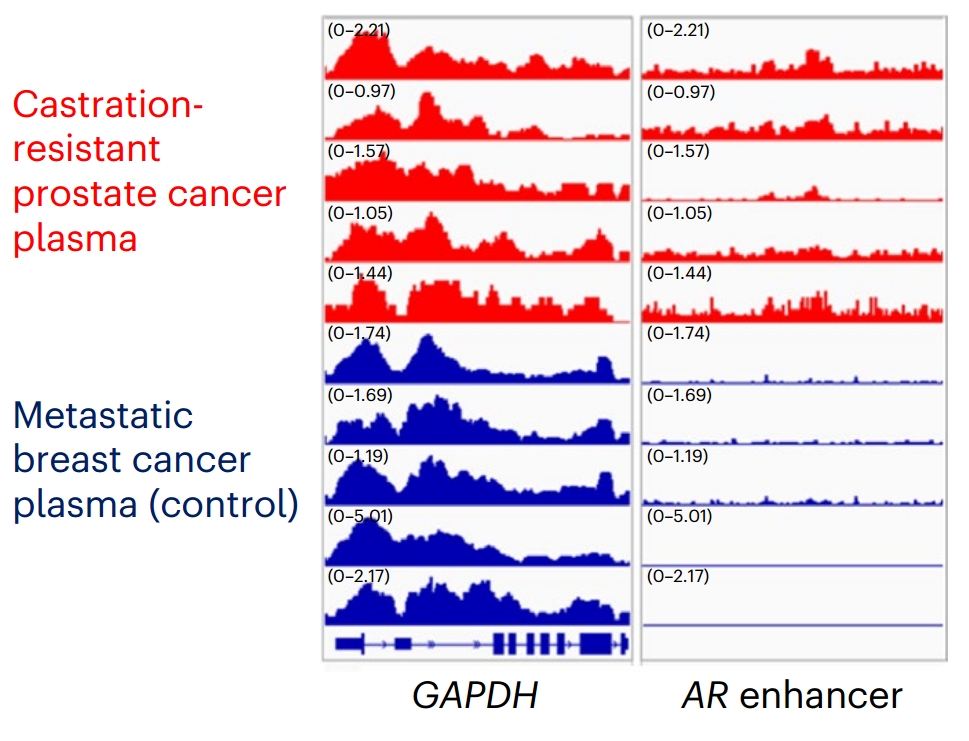
Figure 7. IGV peak diagram of H3K27ac signal in androgen receptor gene enhancer in plasma samples of prostate cancer patients
References
[1] Sadeh R, Sharkia I, Fialkoff G, et al. ChIP-seq of plasma cell-free nucleosomes identifies gene expression programs of the cells of origin. Nat Biotechnol. 2021;39(5):586-598. doi:10.1038/s41587-020-00775-6
[2] Baca SC, Seo JH, Davidsohn MP, et al. Liquid biopsy epigenomic profiling for cancer subtyping. Nat Med. 2023;29(11):2737-2741. doi:10.1038/s41591-023-02605-z
